| Dihydropteroate synthetase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
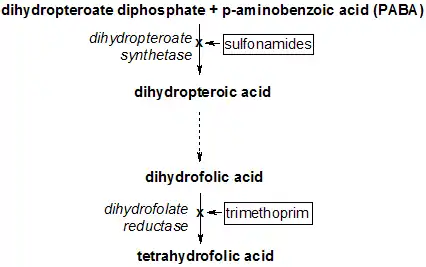 Tetrahydrofolate synthesis pathway | |||||||||
| Identifiers | |||||||||
| EC no. | 2.5.1.15 | ||||||||
| CAS no. | 9055-61-2 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
| Pterin binding enzyme | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||||
| Symbol | Pterin_bind | ||||||||||
| Pfam | PF00809 | ||||||||||
| InterPro | IPR000489 | ||||||||||
| PROSITE | PDOC00630 | ||||||||||
| SCOP2 | 1ajz / SCOPe / SUPFAM | ||||||||||
| |||||||||||
Dihydropteroate synthase is an enzyme classified under EC 2.5.1.15. It produces dihydropteroate in bacteria, but it is not expressed in most eukaryotes including humans. This makes it a useful target for sulfonamide antibiotics, which compete with the PABA precursor.
- (2-amino-4-hydroxy-7,8-dihydropteridin-6-yl)methyl diphosphate + 4-aminobenzoate (PABA) diphosphate + dihydropteroate.
All organisms require reduced folate cofactors for the synthesis of a variety of metabolites. Most microorganisms must synthesize folate de novo because they lack the active transport system of higher vertebrate cells that allows these organisms to use dietary folates. Proteins containing this domain include dihydropteroate synthase (EC 2.5.1.15) as well as a group of methyltransferase enzymes including methyltetrahydrofolate, corrinoid iron-sulphur protein methyltransferase (MeTr)[1] that catalyses a key step in the Wood-Ljungdahl pathway of carbon dioxide fixation.
Dihydropteroate synthase (EC 2.5.1.15) (DHPS) catalyses the condensation of 6-hydroxymethyl-7,8-dihydropteridine pyrophosphate to para-aminobenzoic acid to form 7,8-dihydropteroate. This is the second step in the three-step pathway leading from 6-hydroxymethyl-7,8-dihydropterin to 7,8-dihydrofolate. DHPS is the target of sulfonamides, which are substrate analogues that compete with para-aminobenzoic acid. Bacterial DHPS (gene sul or folP)[2] is a protein of about 275 to 315 amino acid residues that is either chromosomally encoded or found on various antibiotic resistance plasmids. In the fungus Pneumocystis jirovecii (previously P. carinii) DHPS is the C-terminal domain of a multifunctional folate synthesis enzyme (gene fas).[3]
References
- ↑ Universal protein resource accession number Q46389 at UniProt.
- ↑ Crawford IP, Slock J, Stahly DP, Six EW, Han CY (1990). "An apparent Bacillus subtilis folic acid biosynthetic operon containing pab, an amphibolic trpG gene, a third gene required for synthesis of para-aminobenzoic acid, and the dihydropteroate synthase gene". J. Bacteriol. 172 (12): 7211–7226. doi:10.1128/jb.172.12.7211-7226.1990. PMC 210846. PMID 2123867.
- ↑ Volpe F, Dyer M, Scaife JG, Darby G, Stammers DK, Delves CJ (1992). "The multifunctional folic acid synthesis fas gene of Pneumocystis carinii appears to encode dihydropteroate synthase and hydroxymethyldihydropterin pyrophosphokinase". Gene. 112 (2): 213–218. doi:10.1016/0378-1119(92)90378-3. PMID 1313386.
External links
- Dihydropteroate+synthetase at the U.S. National Library of Medicine Medical Subject Headings (MeSH)