| signal recognition particle 9kDa | |||||||
|---|---|---|---|---|---|---|---|
| Identifiers | |||||||
| Symbol | SRP9 | ||||||
| NCBI gene | 6726 | ||||||
| HGNC | 11304 | ||||||
| OMIM | 600707 | ||||||
| RefSeq | NM_003133 | ||||||
| UniProt | P49458 | ||||||
| Other data | |||||||
| Locus | Chr. 1 q42.12 | ||||||
| |||||||
| signal recognition particle 14kDa | |||||||
|---|---|---|---|---|---|---|---|
| Identifiers | |||||||
| Symbol | SRP14 | ||||||
| NCBI gene | 6727 | ||||||
| HGNC | 11299 | ||||||
| OMIM | 600708 | ||||||
| RefSeq | NM_003134 | ||||||
| UniProt | P37108 | ||||||
| Other data | |||||||
| Locus | Chr. 15 q22 | ||||||
| |||||||
| signal recognition particle 19kDa | |||||||
|---|---|---|---|---|---|---|---|
| Identifiers | |||||||
| Symbol | SRP19 | ||||||
| NCBI gene | 6728 | ||||||
| HGNC | 11300 | ||||||
| OMIM | 182175 | ||||||
| RefSeq | NM_003135 | ||||||
| UniProt | P09132 | ||||||
| Other data | |||||||
| Locus | Chr. 5 q21-q22 | ||||||
| |||||||
| signal recognition particle 54kDa | |||||||
|---|---|---|---|---|---|---|---|
| Identifiers | |||||||
| Symbol | SRP54 | ||||||
| NCBI gene | 6729 | ||||||
| HGNC | 11301 | ||||||
| OMIM | 604857 | ||||||
| RefSeq | NM_003136 | ||||||
| UniProt | P61011 | ||||||
| Other data | |||||||
| Locus | Chr. 14 q13.2 | ||||||
| |||||||
| signal recognition particle 68kDa | |||||||
|---|---|---|---|---|---|---|---|
| Identifiers | |||||||
| Symbol | SRP68 | ||||||
| NCBI gene | 6730 | ||||||
| HGNC | 11302 | ||||||
| OMIM | 604858 | ||||||
| RefSeq | NM_014230 | ||||||
| UniProt | Q9UHB9 | ||||||
| Other data | |||||||
| Locus | Chr. 17 q25.1 | ||||||
| |||||||
| signal recognition particle 72kDa | |||||||
|---|---|---|---|---|---|---|---|
| Identifiers | |||||||
| Symbol | SRP72 | ||||||
| NCBI gene | 6731 | ||||||
| HGNC | 11303 | ||||||
| OMIM | 602122 | ||||||
| RefSeq | NM_006947 | ||||||
| UniProt | O76094 | ||||||
| Other data | |||||||
| Locus | Chr. 4 q11 | ||||||
| |||||||
| Signal recognition particle protein | |||||||
|---|---|---|---|---|---|---|---|
| Identifiers | |||||||
| Organism | |||||||
| Symbol | ffh | ||||||
| Alt. symbols | p48, Srp54 | ||||||
| UniProt | P0AGD7 | ||||||
| |||||||
The signal recognition particle (SRP) is an abundant, cytosolic, universally conserved ribonucleoprotein (protein-RNA complex) that recognizes and targets specific proteins to the endoplasmic reticulum in eukaryotes and the plasma membrane in prokaryotes.[1]
History
The function of SRP was discovered by the study of processed and unprocessed secretory proteins, particularly immunoglobulin light chains;[2] and bovine preprolactin. Newly synthesized proteins in eukaryotes carry N-terminal hydrophobic signal sequences, which are bound by SRP when they emerge from the ribosome.[3][4]
Mechanism
In eukaryotes, SRP binds to the signal sequence of a newly synthesized peptide as it emerges from the ribosome.[1] This binding leads to the slowing of protein synthesis known as "elongation arrest", a conserved function of SRP that facilitates the coupling of the protein translation and the protein translocation processes.[5] SRP then targets this entire complex (the ribosome-nascent chain complex) to the protein-conducting channel, also known as the translocon, in the endoplasmic reticulum (ER) membrane. This occurs via the interaction and docking of SRP with its cognate SRP receptor[6] that is located in close proximity to the translocon.
In eukaryotes there are three domains between SRP and its receptor that function in guanosine triphosphate (GTP) binding and hydrolysis. These are located in two related subunits in the SRP receptor (SRα and SRβ)[7] and the SRP protein SRP54 (known as Ffh in bacteria).[8] The coordinated binding of GTP by SRP and the SRP receptor has been shown to be a prerequisite for the successful targeting of SRP to the SRP receptor.[9][10]
Upon docking, the nascent peptide chain is inserted into the translocon channel where it enters into the ER. Protein synthesis resumes as SRP is released from the ribosome.[11][12] The SRP-SRP receptor complex dissociates via GTP hydrolysis and the cycle of SRP-mediated protein translocation continues.[13]
Once inside the ER, the signal sequence is cleaved from the core protein by signal peptidase. Signal sequences are therefore not a part of mature proteins.
Composition and evolution
Despite SRP function being analogous in all organisms, its composition varies greatly. The SRP54-SRP RNA core with GTPase activity is shared in all cellular life, but some subunit polypeptides are specific to eukaryotes.
| Eukaryote | Archaea | Bacteria |
|---|---|---|
| SRP9 SRP14 | No | No |
| SRP19 | Yes | No |
| SRP54 | Yes | Ffh |
| SRP68 SRP72 | No | No |
| 7SL RNA | Yes | 6SL |
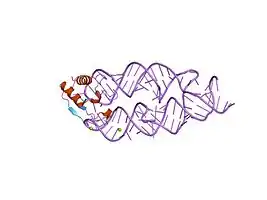 SRP19-7S.S SRP RNA complex from M. jannaschii[14]
SRP19-7S.S SRP RNA complex from M. jannaschii[14]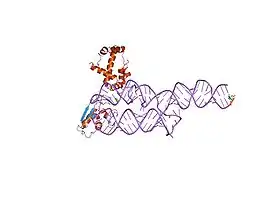 S-domain of human SRP[15]
S-domain of human SRP[15]
Autoantibodies and disease
Anti-signal recognition particle antibodies are mainly associated with, but are not very specific for, polymyositis.[16] For individuals with polymyositis, the presence of anti-SRP antibodies are associated with more prominent muscle weakness and atrophy.[16]
See also
References
- 1 2 Tisdale S, Pellizzoni L (2017-01-01). "Chapter 7 - RNA-Processing Dysfunction in Spinal Muscular Atrophy". In Sumner CJ, Paushkin S, KO CP (eds.). Spinal Muscular Atrophy. Academic Press. pp. 113–131. doi:10.1016/b978-0-12-803685-3.00007-0. ISBN 978-0-12-803685-3.
- ↑ Milstein C, Brownlee GG, Harrison TM, Mathews MB (September 1972). "A possible precursor of immunoglobulin light chains". Nature. 239 (91): 117–20. doi:10.1038/newbio239117a0. PMID 4507519.
- ↑ Walter P, Ibrahimi I, Blobel G (November 1981). "Translocation of proteins across the endoplasmic reticulum. I. Signal recognition protein (SRP) binds to in-vitro-assembled polysomes synthesizing secretory protein". The Journal of Cell Biology. 91 (2 Pt 1): 545–50. doi:10.1083/jcb.91.2.545. PMC 2111968. PMID 7309795.
- ↑ Blobel G, Dobberstein B (December 1975). "Transfer of proteins across membranes. I. Presence of proteolytically processed and unprocessed nascent immunoglobulin light chains on membrane-bound ribosomes of murine myeloma". The Journal of Cell Biology. 67 (3): 835–51. doi:10.1083/jcb.67.3.835. PMC 2111658. PMID 811671.
- ↑ Walter P, Blobel G (December 1983). "Subcellular distribution of signal recognition particle and 7SL-RNA determined with polypeptide-specific antibodies and complementary DNA probe". The Journal of Cell Biology. 97 (6): 1693–9. doi:10.1083/jcb.97.6.1693. PMC 2112735. PMID 6196367.
- ↑ Gilmore R, Blobel G, Walter P (November 1982). "Protein translocation across the endoplasmic reticulum. I. Detection in the microsomal membrane of a receptor for the signal recognition particle". The Journal of Cell Biology. 95 (2 Pt 1): 463–9. doi:10.1083/jcb.95.2.463. PMC 2112970. PMID 6292235.
- ↑ Rapiejko PJ, Gilmore R (May 1992). "Protein translocation across the ER requires a functional GTP binding site in the alpha subunit of the signal recognition particle receptor". The Journal of Cell Biology. 117 (3): 493–503. doi:10.1083/jcb.117.3.493. PMC 2289435. PMID 1315314.
- ↑ Freymann DM, Keenan RJ, Stroud RM, Walter P (January 1997). "Structure of the conserved GTPase domain of the signal recognition particle". Nature. 385 (6614): 361–4. Bibcode:1997Natur.385..361F. doi:10.1038/385361a0. PMID 9002524. S2CID 4238766.
- ↑ Miller JD, Wilhelm H, Gierasch L, Gilmore R, Walter P (November 1993). "GTP binding and hydrolysis by the signal recognition particle during initiation of protein translocation". Nature. 366 (6453): 351–4. Bibcode:1993Natur.366..351M. doi:10.1038/366351a0. PMID 8247130. S2CID 4326097.
- ↑ Grudnik P, Bange G, Sinning I (August 2009). "Protein targeting by the signal recognition particle". Biological Chemistry. 390 (8): 775–82. doi:10.1515/BC.2009.102. PMID 19558326. S2CID 36611716.
- ↑ Lütcke H (March 1995). "Signal recognition particle (SRP), a ubiquitous initiator of protein translocation". European Journal of Biochemistry. 228 (3): 531–50. doi:10.1111/j.1432-1033.1995.0531m.x. PMID 7737147.
- ↑ Luirink J, Sinning I (November 2004). "SRP-mediated protein targeting: structure and function revisited". Biochimica et Biophysica Acta (BBA) - Molecular Cell Research. 1694 (1–3): 17–35. doi:10.1016/j.bbamcr.2004.03.013. PMID 15546655.
- ↑ Shan SO, Walter P (February 2005). "Co-translational protein targeting by the signal recognition particle". FEBS Letters. 579 (4): 921–6. doi:10.1016/j.febslet.2004.11.049. PMID 15680975. S2CID 46046514.
- ↑ Hainzl T, Huang S, Sauer-Eriksson AE (June 2002). "Structure of the SRP19 RNA complex and implications for signal recognition particle assembly". Nature. 417 (6890): 767–71. Bibcode:2002Natur.417..767H. doi:10.1038/nature00768. PMID 12050674. S2CID 2509475.
- ↑ Kuglstatter A, Oubridge C, Nagai K (October 2002). "Induced structural changes of 7SL RNA during the assembly of human signal recognition particle". Nature Structural Biology. 9 (10): 740–4. doi:10.1038/nsb843. PMID 12244299. S2CID 9543041.
- 1 2 Kao AH, Lacomis D, Lucas M, Fertig N, Oddis CV (January 2004). "Anti-signal recognition particle autoantibody in patients with and patients without idiopathic inflammatory myopathy". Arthritis and Rheumatism. 50 (1): 209–15. doi:10.1002/art.11484. PMID 14730618.
Further reading
- Wild K, Becker MM, Kempf G, Sinning I (December 2019). "Structure, dynamics and interactions of large SRP variants". Biological Chemistry. 401 (1): 63–80. doi:10.1515/hsz-2019-0282. PMID 31408431.
- Faoro C, Ataide SF (25 May 2021). "Noncanonical Functions and Cellular Dynamics of the Mammalian Signal Recognition Particle Components". Frontiers in Molecular Biosciences. 8: 679584. doi:10.3389/fmolb.2021.679584. PMC 8185352. PMID 34113652.
External links
- Signal+Recognition+Particle at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- Signal Recognition Particle Database
- www.dnaTube.com video showing an SRP in action
- Another SRP video at www.dnaTube.com
- The Nobel Prize in Physiology or Medicine 1999, "for the discovery that proteins have intrinsic signals that govern their transport and localization in the cell" to Günter Blobel, USA. Press Release, Illustrated Presentation, Presentation Speech