 | |
| Names | |
|---|---|
| IUPAC name
(1S,6R,8R,9R,10S,15R,17R,18R)-8,17-Bis(6-aminopurin-9-yl)-3,12-dihydroxy-3,12-dioxo-2,4,7,11,13,16-hexaoxa-3λ5,12λ5-diphosphatricyclo[13.3.0.06,10]octadecane-9,18-diol | |
| Other names
3',5'-cyclic di-AMP; c-di-AMP; c-di-adenosine monophosphate | |
| Identifiers | |
3D model (JSmol) |
|
| ChEBI | |
| ChemSpider | |
PubChem CID |
|
| UNII | |
| |
| |
| Properties | |
| C20H24N10O12P2 | |
| Molar mass | 658.418 g·mol−1 |
Except where otherwise noted, data are given for materials in their standard state (at 25 °C [77 °F], 100 kPa).
Infobox references | |
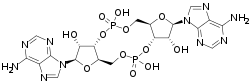
Cyclic di-AMP (also called c-di-AMP and c-di-adenosine monophosphate) is a second messenger used in signal transduction in bacteria and archaea.[1][2][3] It is present in many Gram-positive bacteria, some Gram-negative species, and archaea of the phylum euryarchaeota.[2][3]
It is one of many ubiquitous nucleotide second messengers including cyclic adenosine monophosphate (cAMP), cyclic guanosine monophosphate (cGMP), guanosine pentaphosphate ((p)ppGpp), and cyclic di-GMP (c-di-GMP). c-di-AMP is a signaling nucleotide used in signaling pathways that trigger outputs by using receptor or target proteins to sense c-di-AMP concentrations in the cell.
In bacteria, cyclic di-AMP has been implicated in the control of growth, cell wall homeostasis, bacterial biofilm formation and virulence gene expression, heat and osmotic stress regulation and responses, sporulation, potassium transport, lysis, and antibiotic resistance.[2][4]
In humans, cyclic di-AMP has been implicated in the control of innate immune response and antiviral response against pathogens. The dinucleotide is also produced by numerous human pathogens, prompting the exploration of numerous c-di-AMP-regulating pathways both in humans and in bacteria.
Synthesis
Cyclic di-AMP is synthesized by a membrane-bound diadenylate cyclase (also called diadenylyl cyclase, CdA, and DAC) enzyme called CdaA (DacA). DacA condenses two ATP molecules to make c-di-AMP, releasing 2 pyrophosphates in the process.[5][6] DacA requires a manganese or cobalt metal ion cofactor.[7] Most bacteria possess only one DAC enzyme, but some bacteria like B. subtilis possess two additional DAC enzymes (DisA and CdaS).[2]
Cyclic di-AMP synthesis is inhibited by the GImM I154F mutation in the lactococcus lactis bacterium. GImM is the phosphoglucosamine mutase enzyme that interconverts glucosamine-6-phosphate to glucosamine-1-phosphate to later form cell wall peptidoglycan and other polymers.[4] The I154F mutation inhibits CdA activity by binding to it more strongly than wild-type GImM binds.[4] Thus, GImM modulates c-di-AMP levels.
Synthesis is regulated a number of ways, including negative feedback inhibition and upregulation through a decrease in phosphodiesterase.[2]
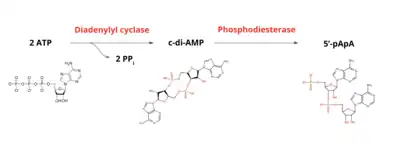 c-di-AMP synthesis and degradation reaction
c-di-AMP synthesis and degradation reaction
Degradation
Phosphodiesterase (PDE) enzymes degrade cyclic di-AMP to the linear molecule 5’-pApA (phosphadenylyl adenosine). 5'-pApA is also involved in a feedback inhibition loop that limits GdpP gene-dependent c-di-AMP hydrolysis, leading to elevated c-di-AMP levels.[8]
Regulation
Since cyclic di-AMP is a signaling nucleotide, it is presumed to adhere to the same regulation pathways, where environmental changes are sensed by synthesis or degradation enzymes, which modulate enzyme concentration. Regulation of c-di-AMP is critical because high c-di-AMP levels lead to abnormal physiology, growth defects, and reduced virulence in infection.[9] In some bacteria, loss of the phosphodiesterases that degrade c-di-AMP leads to cell death.[9][10][11]
It is possible that in addition to enzymatic regulation, intracellular c-di-AMP levels can be regulated by active transport via multidrug resistance transporters that secrete c-di-AMP from the cytoplasm. Listeria monocytogenes has demonstrated such an effect.[9]
At high concentrations, cyclic di-AMP binds to receptor and target proteins to control specific pathways. Elevated c-di-AMP levels have also been linked to increased resistance toward cell wall-damaging antibiotics (e.g. β-lactams) and reduced cellular turgor.[12][13]
Fatty acid synthesis
Cyclic di-AMP has been linked to fatty acid synthesis regulation in Mycobacterium smegmatis, the growth of S. aureus in conditions of low potassium, the sensing of DNA integrity in B. subtilis, and cell wall homeostasis in multiple species.[14][15][16][17]
Cell wall precursor, and thus peptidoglycan precursor, biosynthesis activity can also affect c-di-AMP levels in the cell.[4] Similarly, c-di-AMP levels affect peptidoglycan precursor synthesis, suggesting a strong link between the c-di-AMP and peptidoglycan synthetic pathways.[17]
Cell lysis and RNA synthesis
It is suggested that cyclic di-AMP is involved in the regulation of cell lysis. Studies have shown that bacterial mutant strains with low c-di-AMP levels lysed significantly faster than their parent strains.[4][18]
Cyclic di-AMP has also been linked to bacterial RNA synthesis inhibition. c-di-AMP stimulates the production of (p)ppGpp, an alarmone involved in bacterial stringent response.[19]
STING pathway
In eukaryotic cells, c-di-AMP is sensed and subsequently elicits a type I interferon (IFN) response, leading to the activation of defense mechanisms against viral infection. This detection and activation pathway involves STING, TBK1, and IRF3.[20][21] c-di-AMP may also stimulate dendritic cells, leading to T cell activation.[22]
c-di-AMP activates the innate immune pathway STING (stimulator of interferon genes) to detect damaged DNA. The nucleotide either binds to the helicase DDX41, which in turn activates the STING pathway, or directly binds to the STING protein.[23] Cyclic di-AMP has been identified (along with 2’3’-cGAMP) as a ligand that induces closing of the STING dimer, leading to STING polymerization and pathway activation.[24] When a type I IFN response is not induced in response to c-di-AMP, STING is unable to relocate from the endoplasmic reticulum to the cytoplasm for pathway activation, suggesting that c-di-AMP is a predominant ligand in STING polymerization, and thus activation, via intracellular translocation.[24][25]
See also
References
- ↑ Dey B, Dey RJ, Cheung LS, Pokkali S, Guo H, Lee JH, Bishai WR (April 2015). "A bacterial cyclic dinucleotide activates the cytosolic surveillance pathway and mediates innate resistance to tuberculosis". Nature Medicine. 21 (4): 401–6. doi:10.1038/nm.3813. PMC 4390473. PMID 25730264.
- 1 2 3 4 5 Corrigan RM, Gründling A (August 2013). "Cyclic di-AMP: another second messenger enters the fray". Nature Reviews. Microbiology. 11 (8): 513–24. doi:10.1038/nrmicro3069. hdl:10044/1/19649. PMID 23812326. S2CID 29830881.
- 1 2 Braun F, Thomalla L, van der Does C, Quax TE, Allers T, Kaever V, Albers SV (September 2019). "Cyclic nucleotides in archaea: Cyclic di-AMP in the archaeon Haloferax volcanii and its putative role". MicrobiologyOpen. 8 (9): e00829. doi:10.1002/mbo3.829. PMC 6741144. PMID 30884174.
- 1 2 3 4 5 Zhu Y, Pham TH, Nhiep TH, Vu NM, Marcellin E, Chakrabortti A, et al. (March 2016). "Cyclic-di-AMP synthesis by the diadenylate cyclase CdaA is modulated by the peptidoglycan biosynthesis enzyme GlmM in Lactococcus lactis". Molecular Microbiology. 99 (6): 1015–27. doi:10.1111/mmi.13281. hdl:10453/41116. PMID 26585449. S2CID 24427957.
- ↑ Pham TH, Liang ZX, Marcellin E, Turner MS (November 2016). "Replenishing the cyclic-di-AMP pool: regulation of diadenylate cyclase activity in bacteria". Current Genetics. 62 (4): 731–738. doi:10.1007/s00294-016-0600-8. PMID 27074767. S2CID 1563228.
- ↑ Commichau FM, Heidemann JL, Ficner R, Stülke J (January 2019). Margolin W (ed.). "Making and Breaking of an Essential Poison: the Cyclases and Phosphodiesterases That Produce and Degrade the Essential Second Messenger Cyclic di-AMP in Bacteria". Journal of Bacteriology. 201 (1): e00462–18, /jb/201/1/JB.00462–18.atom. doi:10.1128/JB.00462-18. PMC 6287462. PMID 30224435.
- ↑ Rosenberg J, Dickmanns A, Neumann P, Gunka K, Arens J, Kaever V, et al. (March 2015). "Structural and biochemical analysis of the essential diadenylate cyclase CdaA from Listeria monocytogenes". The Journal of Biological Chemistry. 290 (10): 6596–606. doi:10.1074/jbc.M114.630418. PMC 4358292. PMID 25605729.
- ↑ Bowman L, Zeden MS, Schuster CF, Kaever V, Gründling A (December 2016). "New Insights into the Cyclic Di-adenosine Monophosphate (c-di-AMP) Degradation Pathway and the Requirement of the Cyclic Dinucleotide for Acid Stress Resistance in Staphylococcus aureus". The Journal of Biological Chemistry. 291 (53): 26970–26986. doi:10.1074/jbc.M116.747709. PMC 5207132. PMID 27834680.
- 1 2 3 Huynh TN, Woodward JJ (April 2016). "Too much of a good thing: regulated depletion of c-di-AMP in the bacterial cytoplasm". Current Opinion in Microbiology. Cell regulation. 30: 22–29. doi:10.1016/j.mib.2015.12.007. PMC 4821758. PMID 26773214.
- ↑ Gundlach J, Mehne FM, Herzberg C, Kampf J, Valerius O, Kaever V, Stülke J (October 2015). O'Toole GA (ed.). "An Essential Poison: Synthesis and Degradation of Cyclic Di-AMP in Bacillus subtilis". Journal of Bacteriology. 197 (20): 3265–74. doi:10.1128/JB.00564-15. PMC 4573722. PMID 26240071.
- ↑ Blötz C, Treffon K, Kaever V, Schwede F, Hammer E, Stülke J (2017-07-13). "Mycoplasma pneumoniae". Frontiers in Microbiology. 8: 1328. doi:10.3389/fmicb.2017.01328. PMC 5508000. PMID 28751888.
- ↑ Wang X, Davlieva M, Reyes J, Panesso D, Arias CA, Shamoo Y (March 2017). "A Novel Phosphodiesterase of the GdpP Family Modulates Cyclic di-AMP Levels in Response to Cell Membrane Stress in Daptomycin-Resistant Enterococci". Antimicrobial Agents and Chemotherapy. 61 (3): e01422–16, /aac/61/3/e01422–16.atom. doi:10.1128/AAC.01422-16. PMC 5328519. PMID 28069645.
- ↑ Commichau FM, Gibhardt J, Halbedel S, Gundlach J, Stülke J (March 2018). "A Delicate Connection: c-di-AMP Affects Cell Integrity by Controlling Osmolyte Transport". Trends in Microbiology. 26 (3): 175–185. doi:10.1016/j.tim.2017.09.003. PMID 28965724.
- ↑ Zhang L, Li W, He ZG (February 2013). "DarR, a TetR-like transcriptional factor, is a cyclic di-AMP-responsive repressor in Mycobacterium smegmatis". The Journal of Biological Chemistry. 288 (5): 3085–96. doi:10.1074/jbc.M112.428110. PMC 3561532. PMID 23250743.
- ↑ Corrigan RM, Campeotto I, Jeganathan T, Roelofs KG, Lee VT, Gründling A (May 2013). "Systematic identification of conserved bacterial c-di-AMP receptor proteins". Proceedings of the National Academy of Sciences of the United States of America. 110 (22): 9084–9. Bibcode:2013PNAS..110.9084C. doi:10.1073/pnas.1300595110. PMC 3670340. PMID 23671116.
- ↑ Mehne FM, Gunka K, Eilers H, Herzberg C, Kaever V, Stülke J (January 2013). "Cyclic di-AMP homeostasis in bacillus subtilis: both lack and high level accumulation of the nucleotide are detrimental for cell growth". The Journal of Biological Chemistry. 288 (3): 2004–17. doi:10.1074/jbc.M112.395491. PMC 3548507. PMID 23192352.
- 1 2 Luo Y, Helmann JD (February 2012). "Analysis of the role of Bacillus subtilis σ(M) in β-lactam resistance reveals an essential role for c-di-AMP in peptidoglycan homeostasis". Molecular Microbiology. 83 (3): 623–39. doi:10.1111/j.1365-2958.2011.07953.x. PMC 3306796. PMID 22211522.
- ↑ Witte CE, Whiteley AT, Burke TP, Sauer JD, Portnoy DA, Woodward JJ (May 2013). Mekalanos J (ed.). "Cyclic di-AMP is critical for Listeria monocytogenes growth, cell wall homeostasis, and establishment of infection". mBio. 4 (3): e00282-13. doi:10.1128/mBio.00282-13. PMC 3663569. PMID 23716572.
- ↑ Corrigan RM, Bowman L, Willis AR, Kaever V, Gründling A (February 2015). "Cross-talk between two nucleotide-signaling pathways in Staphylococcus aureus". The Journal of Biological Chemistry. 290 (9): 5826–39. doi:10.1074/jbc.M114.598300. PMC 4342491. PMID 25575594.
- ↑ Jin L, Hill KK, Filak H, Mogan J, Knowles H, Zhang B, et al. (September 2011). "MPYS is required for IFN response factor 3 activation and type I IFN production in the response of cultured phagocytes to bacterial second messengers cyclic-di-AMP and cyclic-di-GMP". Journal of Immunology. 187 (5): 2595–601. doi:10.4049/jimmunol.1100088. PMC 3159690. PMID 21813776.
- ↑ Burdette DL, Monroe KM, Sotelo-Troha K, Iwig JS, Eckert B, Hyodo M, et al. (September 2011). "STING is a direct innate immune sensor of cyclic di-GMP". Nature. 478 (7370): 515–8. Bibcode:2011Natur.478..515B. doi:10.1038/nature10429. PMC 3203314. PMID 21947006.
- ↑ Škrnjug I, Rueckert C, Libanova R, Lienenklaus S, Weiss S, Guzmán CA (2014-04-22). "The mucosal adjuvant cyclic di-AMP exerts immune stimulatory effects on dendritic cells and macrophages". PLOS ONE. 9 (4): e95728. Bibcode:2014PLoSO...995728S. doi:10.1371/journal.pone.0095728. PMC 3996008. PMID 24755640.
- ↑ Parvatiyar K, Zhang Z, Teles RM, Ouyang S, Jiang Y, Iyer SS, et al. (December 2012). "The helicase DDX41 recognizes the bacterial secondary messengers cyclic di-GMP and cyclic di-AMP to activate a type I interferon immune response". Nature Immunology. 13 (12): 1155–61. doi:10.1038/ni.2460. PMC 3501571. PMID 23142775.
- 1 2 Ergun SL, Fernandez D, Weiss TM, Li L (July 2019). "STING Polymer Structure Reveals Mechanisms for Activation, Hyperactivation, and Inhibition". Cell. 178 (2): 290–301.e10. doi:10.1016/j.cell.2019.05.036. PMID 31230712.
- ↑ Barker JR, Koestler BJ, Carpenter VK, Burdette DL, Waters CM, Vance RE, Valdivia RH (April 2013). Taylor R (ed.). "STING-dependent recognition of cyclic di-AMP mediates type I interferon responses during Chlamydia trachomatis infection". mBio. 4 (3): e00018-13. doi:10.1128/mBio.00018-13. PMC 3663186. PMID 23631912.