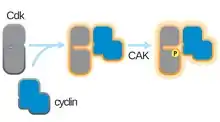
A cyclin-dependent kinase complex (CDKC, cyclin-CDK) is a protein complex formed by the association of an inactive catalytic subunit of a protein kinase, cyclin-dependent kinase (CDK), with a regulatory subunit, cyclin.[1] Once cyclin-dependent kinases bind to cyclin, the formed complex is in an activated state. Substrate specificity of the activated complex is mainly established by the associated cyclin within the complex. Activity of CDKCs is controlled by phosphorylation of target proteins, as well as binding of inhibitory proteins.[2]
Structure and Regulation
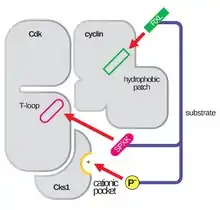
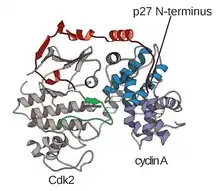
The structure of CDKs in complex with a cyclin subunits (CDKC) has long been a goal of structural and cellular biologists starting in the 1990s when the structure of unbound cyclin A was solved by Brown et al. and in the same year Jeffery et al. solved the structure of human cyclin A-CDK2 complex to 2.3 Angstrom resolution.[3] Since this time, many CDK structures have been determined to higher resolution, including the structures of CDK2 and CDK2 bound to a variety of substrates, as seen in Figure 1. High resolution structures exist for approximately 25 CDK-cyclin complexes in total within the Protein Data Bank.[4] Based on function, there are two general populations of CDK-cyclin complex structures, open and closed form. The difference between the forms lies within the binding of cyclin partners where closed form complexes have CDK-cyclin binding at both the C and N-termini of the activation loop of the CDK, whereas the open form partners bind only at the N-terminus. Open form structures correspond most often to those complexes involved in transcriptional regulation (CDK 8, 9, 12, and 13), while closed form CDK-cyclin complex are most often involved in cell cycle progression and regulation (CDK 1, 2, 6). These distinct roles, however, do not significantly differ with the sequence homology between the CDK components. In particular, among these known structures there appear to be four major conserved regions: a N-terminal Glycine-rich loop, a Hinge Region, an αC-helix, and a T-loop regulation site.[4]
Activation Loop
The activation loop, also referred to as the T-loop, is the region of CDK (between the DFG and APE motifs in many CDK)[4] that is enzymatically active when CDK is bound to its function-specific partner. In CDK-cyclin complexes, this activation region is composed of a conserved αL-12 Helix and contains a key phosphorylatable residue (usually Threonine for CDK-cyclin partners, but also includes Serine and Tyrosine) that mediates the enzymatic activity of the CDK. It is at this essential residue (T160 in CDK2 complexes, T177 in CDK6 complexes) that enzymatic ATP-phosphorylation of CDK-cyclin complexes by CAK (cyclin activating kinase, referring to the CDK7-Cyclin H complex in human cells) takes place. After the hydrolysis of ATP to phosphorylate at this site, these complexes are able to complete their intended function, the phosphorylation of cellular targets. It is important to note that in CDK 1, 2 and 6, the T-loop and a separate C-terminal region are the major sites of cyclin binding in the CDK, and which cyclins are bound to each of these CDK is mediated by the particular sequence of the activation site T-loop. These cyclin binding sites are the regions of highest variability in CDKs despite relatively high sequence homology surrounding the αL-12 Helix motif of this structural component.[4]
Glycine-rich region
The glycine-rich loop (Gly-rich loop) as seen in residues 12-16 in CDK2 encodes a conserved GXGXXG motif across both yeast and animal models. The regulatory region is subject to differential phosphorylation at non-glycine residues within this motif, making this site subject to Wee1 and/or Myt1 inhibitory kinase phosphorylation and Cdc25 de-phosphorylation in mammals. This reversible phosphorylation at the Gly-rich loop in CDK2 occurs at Y15, where activity has been further studied. Study of this residue has shown that phosphorylation promotes a conformational change that prevents ATP and substrate binding by steric interference with these necessary binding sites in the activation loop of the CDK-cyclin complexes. This activity is aided by the notable flexibility that the Gly-rich loop has within the structure of most CDK allowing for its rotation toward the activation loop to have a significant effect on reducing substrate affinity without major changes in the overall CDK-cyclin complex structure.[3][5]
Hinge Region
The conserved hinge region of CDK within eukaryotic cells acts as an essential bridge between the Gly-rich loop and the activation loop. CDK are characterized by a N-terminal lobe that is primarily twisted beta-sheet connected via this hinge region to an alpha helix dominated C-terminal lobe. In discussion of the T-loop and the Gly-rich loop, it is important to note that these regions, which must be able to spatially interact in order to carry out their biochemical functions, lie on opposite lobes of the CDK itself. Thus, this hinge region, which can vary in length slightly between CDK type and CDK-cyclin complex, connects essential regulatory regions of the CDK by connecting these lobes, and plays key roles in the resulting structure of CDK-cyclin complexes by properly orienting ATP for easy catalysis of phosphorylation reactions by the assembled complex. [3][4]
αC-Helix
The αC-Helix region is highly conserved across many of the mammalian kinome (family of kinases). Its main responsibility is to maintain allosteric control of the kinase active site. This control manifests in CDK-cyclin complexes by specifically preventing CDK activity until its binds to its partner regulator (i.e. cyclin or other partner protein). This binding causes a conformational change in the αC-Helix region of the CDK and allows for it to be moved from the active site cleft and completes the initial process of T-loop activation. Given that this region is so conserved across the protein superfamily of kinases, this mechanism where the αC-Helix has been shown to fold out of the N-terminal lobe of the kinase, allowing for increased access to the αL-12 Helix that lies within the T-loop, is considered a potential target for drug development.[6]
The cell cycle
Yeast cell cycle
Although these complexes have a variety functions, CDKCs are most known for their role in the cell cycle. Initially, studies were conducted in Schizosaccharomyces pombe and Saccharomyces cerevisiae (yeast). S. pombe and S. cerevisiae are most known for their association with a single Cdk, Cdc2 and Cdc28 respectively, which complexes with several different cyclins.[7] Depending on the cyclin, various portions of the cell cycle are affected. For example, in S. pombe, Cdc2 associates with Cdk13 to form the Cdk13-Cdc2 complex. In S. cerevisiae, the association of Cdc28 with cyclins, Cln1, Cln2, or Cln3, results in the transition from G1 phase to S phase. Once in the S phase, Cln1 and Cln2 dissociates with Cdc28 and complexes between Cdc28 and Clb5 or Clb6 are formed. In G2 phase, complexes formed from the association between Cdc28 and Clb1, Clb2, Clb3, or Clb4, results in the progression from G2 phase to M (Mitotic) phase. These complexes are present in early M phase as well.[2] See Table 1 for a summary of yeast CDKCs.
- Table 1. CDKCs Associated with Cell Cycle Phases in Yeast
| CDK | Cyclin | Cell Cycle Phase |
|---|---|---|
| Cdc2 (S. pombe) | Cdc13 | G2 to M phase transition; early M phase |
| Cdc28 (S. cerevisiae) | Cln1, Cln2 | G1 to S phase transition |
| Cdc28 | Clb5, Clb6 | S phase |
| Cdc28 | Clb1, Clb2, Clb3, Clb4 | G2 to M phase transition; early M phase |
From what is known about the complexes formed during each phase of the cell cycle in yeast, proposed models have emerged based on important phosphorylation sites and transcription factors involved.[7][8]
Mammalian cell cycle
Using the information discovered through yeast cell cycle studies, significant progress has been made regarding the mammalian cell cycle. It has been determined that the cell cycles are similar and CDKCs, either directly or indirectly, affect the progression of the cell cycle. As previously mentioned, in yeast, only one cyclin-dependent kinase (CDK) is associated with several different cyclins. However, in mammalian cells, several different CDKs bind to various cyclins to form CDKCs. For instance, Cdk1 (also known as human Cdc2), the first human CDK to be identified, associates with cyclins A or B. CyclinA/B-Cdk1 complexes drive the transition between G2 phase and M phase, as well as early M phase. Another mammalian CDK, Cdk2, can form complexes with cyclins D1, D2, D3, E, or A. Cdk4 and Cdk6 interact with cyclins D1, D2, and D3.[9] Studies have indicated that there is no difference between CDKCs cyclin D1-Cdk4/6, therefore, any unique properties can possibly be linked to substrate specificity or activation.[1] While levels of CDKs remain fairly constant throughout the cell cycle, cyclin levels fluctuate. The fluctuation controls the activation of the cyclin-CDK complexes and ultimately the progression throughout the cycle.[10] See Table 2 for a summary of mammalian cell CDKCs involved in the cell cycle.
- Table 2. CDKCs Associated with Cell Cycle Phases in Mammalian Cells[4]
| CDK | Cyclin | Cell Cycle Phase | Non-Cyclin Partner Proteins |
|---|---|---|---|
| Cdk1 (Cdc2) | Cyclins A and B | G2 to M phase transition; early M phase | Cks1 and Cks2 |
| Cdk2 | Cyclins D1, D2, D3 | G1 phase | KAP, Cks1, p27KIP1, and Spy-1 |
| Cdk2 | Cyclin E | G1 to S phase transition | KAP, Cks1, p27KIP1, and Spy-1 |
| Cdk2 | Cyclin A | S phase | KAP, Cks1, p27KIP1, and Spy-1 |
| Cdk4 | Cyclins D1, D2, D3 | G1 phase | HSP90-Cdc37 |
| Cdk6 | Cyclins D1, D2, D3 | G1 phase | p16INK4A, p19INK4D, and P18INK4C-cyclin K |
| Cdk8 | Cyclin C | --- | --- |
| Cdk9 | Cyclin T | --- | Tat, AFF4, and TAR |
| Cdk12 | Cyclin K | --- | --- |
| Cdk13 | Cyclin K | --- | --- |
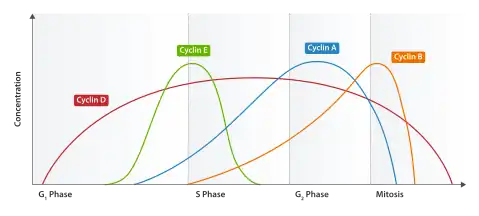
G1 to S phase progression
During late G1 phase, CDKCs bind and phosphorylate members of the retinoblastoma (Rb) protein family. Members of the Rb protein family are tumor suppressors, which prevent uncontrolled cell proliferation that would occur during tumor formation. However, pRbs are also thought to repress the genes required in order for the transition from G1 phase to S phase to occur. When the cell is ready to transition into the next phase, CDKCs, cyclin D1-Cdk4 and cyclin D1-Cdk6 phosphorylate pRB, followed by additional phosphorylation from the cyclin E-Cdk2 CDKC.[11][12] Once phosphorylation occurs, transcription factors are then released to irreversibly inactivate pRB and progression into the S phase of the cell cycle ensues.[13] The cyclin E-Cdk2 CDKC formed in the G1 phase then aids in the initiation of DNA replication during S phase.[1]
G2 to M phase progression
At the end of S phase, cyclin A is associated with Cdk1 and Cdk2. During G2 phase, cyclin A is degraded, while cyclin B is synthesized and cyclin B-Cdk1 complexes form. Not only are cyclin B-Cdk1 complexes important for the transition into M phase, but these CDKCs play a role in the following regulatory and structural processes:[1]
- Chromosomal condensation
- Fragmentation of Golgi network
- Breakdown of nuclear lamina
Inactivation of the cyclin B-Cdk1 complex through the degradation of cyclin B is necessary for exit out of the M phase of the cell cycle.[1]
Other
Even though the majority of the known CDKCs are involved in the cell cycle, not all kinase complexes function in this manner. Studies have shown other CDKCs, such as cyclin k-Cdk9 and cyclin T1-Cdk9, are involved in the replication stress response,[14] and influence transcription.[15][16][17] Additionally, cyclin H-Cdk7 complexes may play a role in meiosis in male germ cells,[18] and has been shown to be involved in transcriptional activities as well.[1][19]
See also
References
- 1 2 3 4 5 6 Malumbres M, Barbacid M. Mammalian cyclin-dependent kinases. Trends Biochem. Sci. 2005 Nov;30(11):630-41
- 1 2 Lodish H, Baltimore D, Berk A, Zipursky SL, Matsudaira P, Darnell J. 1995. Molecular Cell Biology. 3rd Ed. New York: Scientific American Books
- 1 2 3 Kristi Levine, Frederick R Cross, Structuring cell-cycle biology, Structure, Volume 3, Issue 11, 1995, Pages 1131-1134, ISSN 0969-2126, doi:10.1016/S0969-2126(01)00248-9.
- 1 2 3 4 5 6 Wood, D. J., & Endicott, J. A. (2018). Structural insights into the functional diversity of the CDK-cyclin family. Open biology, 8(9), 180112.
- ↑ Malumbres: Cyclin-dependent kinases. Genome Biology, 2014, 15:22, doi:10.1186/gb4184
- ↑ Lorenzo Palmieri, Giulio Rastelli, αC helix displacement as a general approach for allosteric modulation of protein kinases, Drug Discovery Today, Volume 18, Issues 7–8, 2013, Pages 407-414, ISSN 1359-6446, doi:10.1016/j.drudis.2012.11.009.
- 1 2 Simon I, Barnett J, Hannett N, Harbison CT, Rinaldi NJ, Volkert TL, Wyrick JJ, Zeitlinger J, Gifford DK, Jaakkola TS, Young RA. Serial regulation of transcriptional regulators in the yeast cell cycle. Cell. 2001 Sep 21;106(6):697-708.
- ↑ Barik D, Baumann WT, Paul MR, Novak B, Tyson JJ. A model of yeast cell-cycle regulation based on multisite phosphorylation. Mol Syst Biol. 2010 Aug 24;6:405.
- ↑ Malumbres M, Barbacid M. Cell cycle, CDKs and cancer: a changing paradigm. Nat Rev Cancer. 2009 Mar;9(3):153-66.
- ↑ Vermeulen K, Van Bockstaele DR, Berneman ZN. The cell cycle: a review of regulation, deregulation and therapeutic targets in cancer. Cell Prolif. 2003 Jun;36(3):131-49.
- ↑ Mittnacht S. Control of pRB phosphorylation. Curr Opin Genet Dev. 1998 Feb;8(1):21-7.
- ↑ Kaelin WG Jr. Functions of the retinoblastoma protein. Bioessays. 1999 Nov;21(11):950-8.
- ↑ Lundberg AS, Weinberg RA. Functional inactivation of the retinoblastoma protein requires sequential modification by at least two distinct cyclin-cdk complexes. Mol Cell Biol. 1998 Feb;18(2):753-61.
- ↑ Yu DS, Zhao R, Hsu EL, Cayer J, Ye F, Guo Y, Shyr Y, Cortez D. Cyclin-dependent kinase 9-cyclin K functions in the replication stress response. EMBO Rep. 2010 Nov;11(11):876-82.
- ↑ Fu TJ, Peng J, Lee G, Price DH, Flores O. Cyclin K functions as a CDK9 regulatory subunit and participates in RNA polymerase II transcription. J Biol Chem. 1999 Dec 3;274(49):34527-30.
- ↑ Yang Z, Zhu Q, Luo K, Zhou Q. The 7SK small nuclear RNA inhibits the CDK9/cyclin T1 kinase to control transcription. Nature. 2001 Nov 15;414(6861):317-22.
- ↑ Yu DS, Cortez D. A role for CDK9-cyclin K in maintaining genome integrity. Cell Cycle. 2011 Jan 1;10(1):28-32.
- ↑ Kim JM, McGaughy JT, Bogle RK, Ravnik SE. Meiotic expression of the cyclin H/Cdk7 complex in male germ cells of the mouse. Biol Reprod. 2001 May;64(5):1400-8.
- ↑ Patel SA, Simon MC. Functional analysis of the Cdk7.cyclin H.Mat1 complex in mouse embryonic stem cells and embryos. J Biol Chem. 2010 May 14;285(20):15587-98.