| Lily virus X | |
|---|---|
| Virus classification | |
| (unranked): | Virus |
| Realm: | Riboviria |
| Kingdom: | Orthornavirae |
| Phylum: | Kitrinoviricota |
| Class: | Alsuviricetes |
| Order: | Tymovirales |
| Family: | Alphaflexiviridae |
| Genus: | Potexvirus |
| Species: | Lily virus X |
Lily virus X (LVX) is a pathogenic ssRNA(+) plant virus of the family Alphaflexiviridae and the order Tymovirales.
Description
LVX is described in the 4th report of the ICTV (1982). It is found primarily in lilies, although more plants are susceptible, and the virus is thought to be only transmitted mechanically. There are no known vectors; however, the application of insecticides effectively reduced the spread of LVX, from which it may be gathered that the virus’ transmission is insect-mediated.[1] Symptoms of this virus are as of yet unknown, obfuscating the capability to examine the extent of natural infection and spread.
Structure

LVX is a non-enveloped virus with helical symmetry. All potexviruses, including LVX are believed to have slightly less than 9 protein subunits per helical turn.[3] This pattern of nucleocapsid formation causes the nucleocapsid to be an elongated, flexible, filamentous virus like most plant viruses. Unlike other potexviruses (average length of 550 nm), LVX has a length of 470 nm and a width of 13 nm.[4] LVX is distinguishable from lily symptomless carlavirus (LSV) by serological tests, such as immunogold labeling tests.[5]
Genome
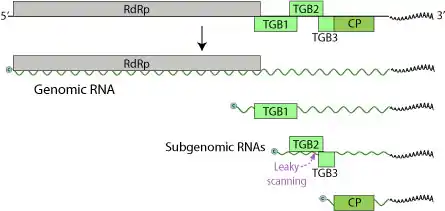
The LVX genome consists of one single stranded (+)RNA sequence of 5.9-7 kilobases in size. The genome contains only three Open Reading Frames (ORFs), coding for five proteins (RdRp, TGBp1, TGBp2, TGBp3, CP).[6] The 5’ end is capped and the 3’ terminus is polyadenylated.
These proteins are 24 kilodaltons (kDa), 12 kDa, and 22 kDa, with the third being the coat protein, while also encoding for RNA-dependent RNA polymerase (RdRp).[7] The stop codon of the 24 kDa ORF and the start codon of the 12 kDa ORF overlap one another. While many potexvirus genomes contain a small ORF that is immediately 5’ of this coat protein ORF, this is not found in the LVX genome. This difference is surprising as this ORF has been identified in at least six other potexvirus genomes. However, when looking at the proteins encoded by LVX and other potexviruses, there were significant similarities between the amino acid sequences. At the 5’ end of LVX, there is a truncated ORF, which encodes for a polypeptide which contains a GDD motif. This motif is also found in the C-terminal end of proteins encoded by other potexviruses. Examination of the 24 kDa protein of LVX reveals that it contains a nucleoside triphosphate binding motif (GXXGXGKS/T). This motif also found in the lily symptomless virus 25 kDa protein, as well as the 25-26 kDa proteins of other potexviruses, such as potato virus X and potato virus S.
LVX also contains a supposed potexvirus subgenomic promoter sequence (GGTTAAGTT---GAA) upstream (5’) of the 24 kDa protein. This sequence is also found upstream of initiation codons of coat proteins in similar viruses, most notably lily symptomless virus. Researchers looked at LVX-infected lily leaves and found the presence of subgenomic RNAs. These RNAs were about 2000 and 850 nucleotides in length, which matched up with the presumed subgenomic promoters. From this, it was determined that they likely function as messengers for both the 24 kDa and 22 kDa proteins (coat protein). In the LVX sequence, there is a region between the 12 kDa ORF and the 22 kDa (coat protein) ORF that is not translated. In other potexvirus sequences, this region contains respective 7 kDa and 11 kDa ORFs; however, in this region of the LVX sequence, there is an uninterrupted reading frame. This reading frame begins at nucleotide position 1236 and ends at position 1646. If this region were to be translated, the protein encoded would be similar to the 7 kDa and 11 kDa proteins mentioned above, ending 18 nucleotides inside of LVX's coat protein ORF. Translation of this LVX ORF does not get translated because the genome is missing an initiation codon.
Triple Gene Block 1 (TGBp1) is a multifunctional protein. It acts to promote translation of viral RNAs by acting as an RNA helicase, separating double-stranded RNA for RdRp functions. Moreover, it can act as a suppressor of RNA interference, which is an immune defense against the accumulation of viral RNAs.[8] TGBp1 transports the viral genome to adjacent plant cells directly through plasmosdesmata, allowing for efficient propagation by circumventing the host's cell wall barrier.[9] TGBp1 also increases plasmodesmal size exclusion limits, allowing the viral genome to pass more easily from one cell to another. Finally, it suppresses RNA silencing, preventing the host from disabling the RNA genome.
TGBp2 and TGBp3 are membrane binding proteins involved in attachment and entry. TGBp3 is expressed through leaky scanning of the TGBp2 subgenomic mRNA. The TGBp2 ORF has a weak initiation codon that is sometimes skipped by the ribosome in translation initiation. In this instance, the 40S ribosomal subunit continues to scan until it encounters the initiation codon of TGBp3 and begins translation.
CP is the coat protein of LVX that forms ribonucleoprotein complexes along with TGBp1 and viral mRNA.
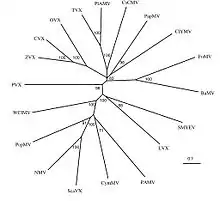
“The nucleotide sequence of LVX appears to be unique among potexviruses inasmuch as it apparently lacks the small open reading frame, 5’ to the coat protein cistron, common to all other potexvirus.”[10] In addition, the TGBp3 region of the genome lacked a normal start codon.[11] The 5′-non-coding region begins with GGAAAA, whereas those of other sequenced potexviruses probably all begin with GAAAA.[12] Phylogenetic analysis of the LVX coding sequence revealed that LVX was most closely related to Strawberry mild yellow-edge virus.[13]
Replication
LVX does not have a known vector, but it most likely spreads and enters the cell through mechanical inoculation by insects. The replication of LVX, like other ssRNA(+) viruses, occurs in the cytoplasm of cells. Once the virus enters into the host cell, the virus is uncoated and releases the viral genome RNA into the cytoplasm. The viral monocistronic RNA is then translated into RNA-dependent RNA polymerase, encoded by the 5’-proximal ORF. The replication of LVX occurs in viral factories which are organized by the protein TGBp1.[15] TGBp1 works to rearrange the actin and endomembranes of the host and creates an assembly of helical arrangements. These helical structures are surrounded by the host endomembranes, which create a region where the virus can replicate more efficiently.[16]
The replication of the ssRNA(+) virus generates a dsRNA(+) virus, which is then further transcribed and replicated to create more LVX mRNAs and ssRNA(+) genomes, respectively. Subgenomic promoters on the virus lead to translation of sgRNAs, which results in the formation of capsid and movement proteins. These are used to arrange the virion structure and organize the viral genomes. New virus particles are then able to be produced and assembled. With the help of LVX's triple gene block proteins, these completed particles can traverse long distances among plasmodesmata and shorter, intercellular spaces in order to infect other host cells.[17]
Associated Disease
LVX has no known associated diseases.
Symptoms
There are no known outstanding symptoms that arise in LVX's natural host range of Lilium formosanum; however, Yang (1997) found that LVX-infected lilies grow more slowly and tend to be smaller than uninfected specimens. Mechanical inoculation of other susceptible plant species, such as Tetragonia tetragonioides and Chenopodium murale, result in chlorotic local lesions.[18]
Tropism
Although not seen in nature, lab tests have shown that LVX is able to infect several other plants beyond Lilium formosanum.[19] Other species that have been shown to be susceptible are:[20]
- Chenopodium capitatum
- Chenopodium murale
- Chenopodium quinoa
- Gomphrena globosa
- Lilium formosanum
- Nicotiana benthamiana
- Nicotiana clevelandii
- Tetragonia tetragonioides
LVX is able to infect all tissues of the lily plant. Virions have been cytopathologically detected in all parts of the host plants and contain approximately 5% nucleic acid and 95% protein, with no lipid contents.[21] The mode of transmission is expected to be that of mechanical inoculation through insect vectors, since the spread of the virus was inhibited by insecticides but not mineral-oil sprays.[22]
References
- ↑ Asjes, C.J. (1991). Control of air-borne field spread of tulip breaking virus, lily symptomless virus and lily virus X in lilies by mineral oils, synthetic pyrethroids, and a nematicide in the Netherlands Netherlands Journal of Plant Path, 97(3), 129-138.
- ↑ Kendall, A., McDonald, M., Bian, W., Bowles, T., Baumgarten, S. C., Shi, J., … Stubbs, G. (2008). Structure of Flexible Filamentous Plant Viruses. Journal of Virology, 82(19), 9546–9554.
- ↑ Kendall, A., McDonald, M., Bian, W., Bowles, T., Baumgarten, S. C., Shi, J., … Stubbs, G. (2008). Structure of Flexible Filamentous Plant Viruses. Journal of Virology, 82(19), 9546–9554.
- ↑ Stone, O.M. (1980). Two new potexviruses from monocotyledons. Acta Hort. 110, 59-63.
- ↑ Yang, T. (1997). Cytological Characteristics and detection of viruses of Lilium Spp. (Doctoral dissertation). University of Florida, Gainesville, Florida.
- ↑ Memelink, J., van der Vlugt, C.I.M., Linthorst, H.J.M., Derks, A.F.L.M., Asjes, C.J., Bol, J.F. (1990). Homologies between the genomes of a carlavirus (lily symptomless virus) and a potexvirus (lily virus X) from lily plants. Journal of General Virology, 71, 917-924.
- ↑ Memelink, J., van der Vlugt, C.I.M., Linthorst, H.J.M., Derks, A.F.L.M., Asjes, C.J., Bol, J.F. (1990). Homologies between the genomes of a carlavirus (lily symptomless virus) and a potexvirus (lily virus X) from lily plants. Journal of General Virology, 71, 917-924.
- ↑ Lubicz-Verchot, J. (2005). A New Cell-to-Cell Transport Model for Potexviruses. Molecular Plant-Microbe Interactions, 18(4), 283-290.
- ↑ Lubicz-Verchot, J. (2005). A New Cell-to-Cell Transport Model for Potexviruses. Molecular Plant-Microbe Interactions, 18(4), 283-290.
- ↑ Yang, T. (1997). Cytological Characteristics and detection of viruses of Lilium Spp. (Doctoral dissertation). University of Florida, Gainesville, Florida.
- ↑ Chen, J., Shi, Y.-H., Adams, M.J., Chen, J.-P. (2005). The complete sequence of the genomic RNA of an isolate of Lily virus X (genus Potexvirus). Archives of Virology, 150(4), 825-832.
- ↑ Chen, J., Shi, Y.-H., Adams, M.J., Chen, J.-P. (2005). The complete sequence of the genomic RNA of an isolate of Lily virus X (genus Potexvirus). Archives of Virology, 150(4), 825-832.
- ↑ Chen, J., Shi, Y.-H., Adams, M.J., Chen, J.-P. (2005). The complete sequence of the genomic RNA of an isolate of Lily virus X (genus Potexvirus). Archives of Virology, 150(4), 825-832.
- ↑ Chen, J., Shi, Y.-H., Adams, M.J., Chen, J.-P. (2005). The complete sequence of the genomic RNA of an isolate of Lily virus X (genus Potexvirus). Archives of Virology, 150(4), 825-832.
- ↑ Yang, T. (1997). Cytological Characteristics and detection of viruses of Lilium Spp. (Doctoral dissertation). University of Florida, Gainesville, Florida.
- ↑ Lubicz-Verchot, J. (2005). A New Cell-to-Cell Transport Model for Potexviruses. Molecular Plant-Microbe Interactions, 18(4), 283-290.
- ↑ Yang, T. (1997). Cytological Characteristics and detection of viruses of Lilium Spp. (Doctoral dissertation). University of Florida, Gainesville, Florida.
- ↑ Stone, O.M. (1980). Two new potexviruses from monocotyledons. Acta Hort. 110, 59-63.
- ↑ Stone, O.M. (1980). Two new potexviruses from monocotyledons. Acta Hort. 110, 59-63.
- ↑ Phillips, S (1986). "Lily X potexvirus". Plant Viruses Online. Archived from the original on December 8, 2015. Retrieved December 7, 2015.
- ↑ Stone, O.M. (1980). Two new potexviruses from monocotyledons. Acta Hort. 110, 59-63.
- ↑ Asjes, C.J. (1991). Control of air-borne field spread of tulip breaking virus, lily symptomless virus and lily virus X in lilies by mineral oils, synthetic pyrethroids, and a nematicide in the Netherlands. Netherlands Journal of Plant Path, 97(3), 129-138.