| Part of a series on |
| Microbial and microbot movement |
|---|
.png.webp) |
| Microswimmers |
| Molecular motors |
|
A microswimmer is a microscopic object with the ability to move in a fluid environment.[1] Natural microswimmers are found everywhere in the natural world as biological microorganisms, such as bacteria, archaea, protists, sperm and microanimals. Since the turn of the millennium there has been increasing interest in manufacturing synthetic and biohybrid microswimmers. Although only two decades have passed since their emergence, they have already shown promise for various biomedical and environmental applications.[1]
Given the recent nature of the field, there is yet no consensus in the literature for the nomenclature of the microscopic objects this article refers to as "microswimmers". Among the many alternative names such objects are given in the literature, microswimmers, micro/nanorobots and micro/nanomotors are likely the most frequently encountered. Other common terms may be more descriptive, including information about the object shape, e.g., microtube or microhelix, its components, e.g., biohybrid, spermbot,[2] bacteriabot,[3] or micro-bio-robot,[4] or behavior, e.g., microrocket, microbullet, microtool or microroller. Researchers have also named their specific microswimmers e.g., medibots,[5] hairbots,[6] iMushbots,[7] IRONSperm,[8] teabots,[9] biobots,[10] T-budbots,[11] or MOFBOTS.[12][1]
Background
In 1828, the British biologist Robert Brown discovered the incessant jiggling motion of pollen in water and described his finding in his article "A Brief Account of Microscopical Observations…",[13] leading to extended scientific discussion about the origin of this motion. This enigma was resolved only in 1905, when Albert Einstein published his celebrated essay Über die von der molekularkinetischen Theorie der Wärme geforderte Bewegung von in ruhenden Flüssigkeiten suspendierten Teilchen.[14] Einstein not only deduced the diffusion of suspended particles in quiescent liquids, but also suggested these findings could be used to determine particle size — in a sense, he was the world's first microrheologist.[15]
Ever since Newton established his equations of motion, the mystery of motion on the microscale has emerged frequently in scientific history, as famously demonstrated by a couple of articles that should be discussed briefly. First, an essential concept, popularized by Osborne Reynolds, is that the relative importance of inertia and viscosity for the motion of a fluid depends on certain details of the system under consideration.[15] The Reynolds number Re, named in his honor, quantifies this comparison as a dimensionless ratio of characteristic inertial and viscous forces:


"Fast or slow, it exactly retraces its trajectory and it's back where it started".[16]
Here, ρ represents the density of the fluid; u is a characteristic velocity of the system (for instance, the velocity of a swimming particle); l is a characteristic length scale (e.g., the swimmer size); and μ is the viscosity of the fluid. Taking the suspending fluid to be water, and using experimentally observed values for u, one can determine that inertia is important for macroscopic swimmers like fish (Re = 100), while viscosity dominates the motion of microscale swimmers like bacteria (Re = 10−4).[15]
The overwhelming importance of viscosity for swimming at the micrometer scale has profound implications for swimming strategy. This has been discussed memorably by E. M. Purcell, who invited the reader into the world of microorganisms and theoretically studied the conditions of their motion.[16] In the first place, propulsion strategies of large scale swimmers often involve imparting momentum to the surrounding fluid in periodic discrete events, such as vortex shedding, and coasting between these events through inertia. This cannot be effective for microscale swimmers like bacteria: due to the large viscous damping, the inertial coasting time of a micron-sized object is on the order of 1 μs. The coasting distance of a microorganism moving at a typical speed is about 0.1 angstroms (Å). Purcell concluded that only forces that are exerted in the present moment on a microscale body contribute to its propulsion, so a constant energy conversion method is essential.[16][15]
Microorganisms have optimized their metabolism for continuous energy production, while purely artificial microswimmers (microrobots) must obtain energy from the environment, since their on-board-storage-capacity is very limited. As a further consequence of the continuous dissipation of energy, biological and artificial microswimmers do not obey the laws of equilibrium statistical physics, and need to be described by non-equilibrium dynamics.[15] Mathematically, Purcell explored the implications of low Reynolds number by taking the Navier-Stokes equation and eliminating the inertial terms:
where is the velocity of the fluid and is the gradient of the pressure. As Purcell noted, the resulting equation — the Stokes equation — contains no explicit time dependence.[16] This has some important consequences for how a suspended body (e.g., a bacterium) can swim through periodic mechanical motions or deformations (e.g., of a flagellum). First, the rate of motion is practically irrelevant for the motion of the microswimmer and of the surrounding fluid: changing the rate of motion will change the scale of the velocities of the fluid and of the microswimmer, but it will not change the pattern of fluid flow. Secondly, reversing the direction of mechanical motion will simply reverse all velocities in the system. These properties of the Stokes equation severely restrict the range of feasible swimming strategies.[16][15]
As a concrete illustration, consider a mathematical scallop that consists of two rigid pieces connected by a hinge. Can the "scallop" swim by periodically opening and closing the hinge? No: regardless of how the cycle of opening and closing depends on time, the scallop will always return to its starting point at the end of the cycle. Here originated the striking quote: "Fast or slow, it exactly retraces its trajectory and it's back where it started".[16] In light of this scallop theorem, Purcell developed approaches concerning how artificial motion at the micro scale can be generated.[15] This paper continues to inspire ongoing scientific discussion; for example, recent work by the Fischer group from the Max Planck Institute for Intelligent Systems experimentally confirmed that the scallop principle is only valid for Newtonian fluids.[17][15]
Types
Different types of microswimmers are powered and actuated in different ways. Swimming strategies for individual microswimmers [3][18][19][20][21][22] as well as swarms of microswimmers [23][24][25][26][27][28] have been examined down through the years. Typically, microswimmers rely either on external power sources, as it is the case for magnetic,[29] optic,[10] or acoustic control,[30] or employ the fuel available in their surroundings, as is the case with biohybrid or catalytic microswimmers. Magnetic and acoustic actuation are typically compatible with in vivo microswimmer manipulation and catalytic microswimmers can be specifically engineered to employ in vivo fuels. The use of optical forces in biological fluids or in vivo is more challenging, but interesting examples have nevertheless been demonstrated.[10]
Often, researchers choose to take inspiration from nature, either for the entire microswimmer design, or for achieving a desired propulsion type. For example, one of the first bioinspired microswimmers consisted of human red blood cells modified with a flagellum-like artificial component made of filaments of magnetic particles bonded via biotin–streptavidin interactions.[31] More recently, biomimetic swimming inspired by worm-like travelling wave features,[32] shrimp locomotion,[33] and bacterial run-and-tumble motion,[34] was demonstrated by using shaped light.[10]
A different nature-inspired approach is the use of biohybrid microswimmers. These comprise a living component and a synthetic one. Biohybrids most often take advantage of the microscale motion of various biological systems and can also make use of other behaviours characterising the living component.[35] For magnetic bioinspired and biohybrid microswimmers, typical model organisms are bacteria, sperm cells and magnetotactic cells.[36] In addition to the use of magnetic forces, actuation of bioinspired microswimmers was also demonstrated using e.g., acoustic excitation [37] or optical forces.[38] Another nature-inspired behavior related to optical forces is that of phototaxis, which can be exploited by e.g., cargo-carrying microorganisms,[39] synthetic microswimmers [40][41][42] or biohybrid microswimmers.[43] A number of recent review papers are focused on explaining or comparing existing propulsion and control strategies used in microswimmer actuation.[44][45][46][47][48] Magnetic actuation is most often included for controlled in vivo guiding, even for microswimmers which rely on a different type of propulsion. In 2020, Koleoso et al. reviewed the use of magnetic small scale robots for biomedical applications and provide details about the various magnetic fields and actuation systems developed for such purposes.[29][1]
Strategies for the fabrication of microswimmers include two-photon polymerisation 3D printing, photolithography, template-assisted electrodeposition, or bonding of a living component to an inanimate one by exploiting different strategies. More recent approaches exploit 4D printing, which is the 3D printing of stimuli-responsive materials.[49][50][51][52] Further functionalization is often required, either to enable a certain type of actuation, e.g., metal coating for magnetic control or thermoplasmonic responses, or as part of the application, if certain characteristics are required for e.g., sensing, cargo transport, controlled interactions with the environment, or biodegradation.[53][54][55][56][1]
Natural microswimmers
.jpg.webp)

Drawing of Chlamydomonas reinhardtii alga in a co-culture with Escherichia coli bacteria [57]
Motile systems have developed in the natural world over time and length scales spanning several orders of magnitude, and have evolved anatomically and physiologically to attain optimal strategies for self-propulsion and overcome the implications of high viscosity forces and Brownian motion, as shown in the diagram on the right.[58][15]
Some of the smallest known natural motile systems are motor proteins, i.e., proteins and protein complexes present in cells that carry out a variety of physiological functions by transducing chemical energy into mechanical energy. These motor proteins are classified as myosins, kinesins, or dyneins. Myosin motors are responsible for muscle contractions and the transport of cargousing actin filaments as tracks. Dynein motors and kinesin motors, on the other hand, use microtubules to transport vesicles across the cell.[59][60] The mechanism these protein motors use to convert chemical energy into movement depends on ATP hydrolysis, which leads to a conformation modification in the globular motor domain, leading to directed motion.[61][62][15]
Apart from motor proteins, enzymes, traditionally recognized for their catalytic functions in biochemical processes, can function as nanoscale machines that convert chemical energy into mechanical action at the molecular dimension. Diffusion of various enzymes (e.g. urease, and catalase), measured by fluorescent correlated spectroscopy (FCS), increases in a substrate-dependent manner.[63][64] Moreover, when enzymes are membrane-bound, their catalytic actions can drive lipid vesicle movement. For instance, lipid vesicles integrated with enzymes such as transmembrane adenosine 5’-triphosphatase, membrane-bound acid phosphatase, or urease exhibit enhanced mobility correlating with the enzymatic turnover rate.[65]
Bacteria can be roughly divided into two fundamentally different groups, gram-positive and gram-negative bacteria, distinguished by the architecture of their cell envelope. In each case the cell envelope is a complex multi-layered structure that protects the cell from its environment. In gram-positive bacteria, the cytoplasmic membrane is only surrounded by a thick cell wall of peptidoglycan. By contrast, the envelope of gram-negative bacteria is more complex and consists (from inside to outside) of the cytoplasmic membrane, a thin layer of peptidoglycan, and an additional outer membrane, also called the lipopolysaccharide layer. Other bacterial cell surface structures range from disorganised slime layers to highly structured capsules. These are made from secreted slimy or sticky polysaccharides or proteins that provide protection for the cells and are in direct contact with the environment. They have other functions, including attachment to solid surfaces. Additionally, protein appendages can be present on the surface: fimbriae and pili can have different lengths and diameters and their functions include adhesion and twitching motility.[66][67][15]
Specifically, for microorganisms that live in aqueous environments, locomotion refers to swimming, and hence the world is full of different classes of swimming microorganisms, such as bacteria, spermatozoa, protozoa, and algae. Bacteria move due to rotation of hair-like filaments called flagella, which are anchored to a protein motor complex on the bacteria cell wall.[15]
The following table, based on Schwarz et al., 2017,[68] lists some examples of natural or biological microswimmers.
| Name | Image | Size (μm2)a | Speed (μm/s)b | Propulsion mechanism | Natural swimming habitat | Sources | |
|---|---|---|---|---|---|---|---|
| bacterial swimmers (prokaryotes) | Escherichia coli | 0.5 × 2 | 30 | Peritrichous bundles | Intestinal flora | [69] | |
| Serratia marcescens | 1 × 2 | 50 | Peritrichous bundles | Respiratory and urinary tracts (parasitic) | [70] | ||
| Salmonella typhimurium | 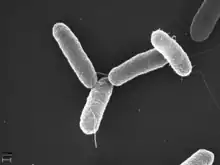 | 0.5 × 2 | 30 | Peritrichous bundles | Intestines (parasitic) | [71] | |
| Bacillus subtilis | 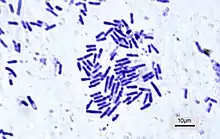 | 1 × 3 | 20 | Peritrichous bundles | Intestinal flora | [72] | |
| Aliivibrio fischeri |  | 1 × 2 | 50 | Lophotrichous flagella | Mucus (symbiotic) | [73] | |
| Vibrio alginolyticus |  | 2 × 3 | 40 | Monotrichous flagellum | Blood (parasitic) | [74][75] | |
| Listeria monocytogenes |  | 0.5 × 1.5 | <1 | Peritrichous or amphitrichous bundles | Inter- and intracellular (parasitic) | [76][77] | |
| Magnetococcus marinus |  | 2 × 2 | 200 | Two lophotrichous bundles | Marine water | [78][79] | |
| Magnetospirillum gryphiswaldense |  | 0.5 × 2 | 60 | Two amphitrichous flagella | Freshwater sediments | [80] | |
| Mycoplasma mobile | 0.5 × 0.5 | 5 | Gliding via protrusions | Fish gills (parasitic) | [81] | ||
| protist swimmers (unicellular) eukaryotes) | Chlamydomonas |  | 10 × 10 | 150 | Two lophotrichous flagella | Freshwater, soil | [82] |
| Tetrahymena |  | 25 × 50 | >500 | Holotrichous cilia | Freshwater | [83] | |
| Trypanosome | 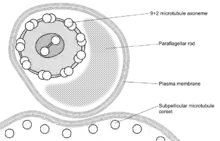 | 3 × 20 | 30 | Monotrichous flagellum | Blood (parasitic) | [84][85] | |
| sperm cells | Human |  | 3 × 5 | 50 | Monotrichous flagellum | Reproductive tract | [86][87] |
| Bovine | 5 × 10 | 100 | Monotrichous flagellum | Reproductive tract | [87][88][89] | ||
| Murine | 3 × 8 | 120 | Monotrichous flagellum | Reproductive tract | [86][88] | ||
Synthetic microswimmers
"An artificial microswimmer is a cutting-edge technology with engineering and medical applications. A natural microswimmer, such as bacteria and sperm cells, also play important roles in wide varieties of engineering, medical and biological phenomena. Due to the small size of the microswimmer, the inertial effect of the surrounding flow field may be negligible. In such a case, reciprocal body deformation cannot induce migration of a swimmer, which is known as the scallop theorem. To overcome the implications of the scallop theorem, the microswimmer needs to undergo a nonreciprocal body deformation to achieve migration. The swimming strategy is thus completely different from macro-scale swimmers...".[90]

One of the current engineering challenges is to create miniaturized functional vehicles that can carry out complex tasks at a small scale that would be otherwise impractical, inefficient, or outright impossible by conventional means. These vehicles are termed nano/micromotors or nano/microrobots, and should be distinguished from even smaller molecular machines for energy, computing, or other applications on the one side and static microelectromechanical systems (MEMS) on the other side of this size scale. Rather than being electronic devices on a chip, micromotors are able to move freely through a liquid medium while being steered or directed externally or by intrinsic design, which can be achieved by various mechanisms, most importantly catalytic reactions,[92][93][94][95] magnetic fields,[96] or ultrasonic waves.[97][98][99][100][101]
There are a variety of sensing, actuating, or pickup-and-delivery applications that scientists are currently aiming for, with local drug targeting for cancer treatment being one of the more prominent examples.[102][5] For applications like this, a micromotor needs to be able to move, i.e., to swim, freely in three dimensions efficiently controlled and directed with a reliable mechanism.[68]
It is a direct consequence of the small size scale of microswimmers that they have a low Reynolds number. This means the physics of how microswimmers swim is dominated by viscous drag forces, a problem which has been discussed extensively by physicists in the field.[99][103][58] This kind of swimming has challenged engineers as it is not commonly experienced in everyday life, but can nonetheless be observed in nature for motile microorganisms like sperm or certain bacteria. Naturally, these microorganisms served as inspiration from the very beginning to create artificial micromotors, as they were able to tackle the challenges that an active, self-sufficient microswimmer vehicle has to face.[104] With biomimetic approaches, researchers were able to imitate the flagella-based motion strategy of sperm and Escherichia coli bacteria by reproducing their respective flagellum shape and actuating it with magnetic fields.[31][105][68][15]
Microorganisms have adapted their locomotion to the harsh environment of low Reynolds number regime by invoking different swimming strategy.[106] For example, the E. coli moves by rotating its helical flagellum,[107][108] Chlamydomonas flagella have a breaststroke kind of motion.[109] African trypanosome has a helical flagellum attached to the cell body with a planar wave passing through it.[110][111] Swimming of these kind of natural swimmers have been investigated for the last half-century.[112] As a result of these studies, artificial swimmers have also been proposed, like Taylor sheet,[113] Purcell's two-hinge swimmer,[16][114] three-linked spheres swimmer,[115][116][117] elastic two-sphere swimmer [118] and three-sphere with a passive elastic arm,[119] which have further enhanced understanding about low Reynolds number swimmers. One of the challenges in proposing an artificial swimmer lies in the fact that the proposed movement stroke should not be reciprocal otherwise it cannot propel itself due to the Scallop theorem. In Scallop theorem, Purcell had argued that a swimmer with one-hinge or one degree of freedom is bound to perform reciprocal motion and thus will not be able to swim in the Stokes regime.[106][16][112]
Purcell proposed two possible ways to elude from Scallop theorem, one is 'corkscrew' motion [107][104] and the other is 'flexible oar' motion.[120][121] Using the concept of flexible oar, Dreyfus et al reported a micro swimmer that exploit elastic property of a slender filament made up of paramagnetic beads.[31] To break the time inversion symmetry, a passive head was attached to the flexible arm. The passive head reduces the velocity of the flexible swimmer, bigger the head, higher is the drag force experienced by the swimmer. The head is essential for swimming because without it the tail performs a reciprocal motion and the velocity of the swimmer reduces to zero.[122][112]
Another way microswimmers can propel is through catalytic reactions. Taking inspiration from Whitesides, who used the decomposition of hydrogen peroxide (H2O2) to propel cm/mm-scale objects on a water surface,[123] Sen et al. (2004) fabricated catalytic motors in the micrometer range.[92] These microswimmers were rod-shaped particles 370 nm in diameter and consisted of 1 µm long Pt and Au segments. They propelled via the decomposition of hydrogen peroxide in solution which would be catalyzed into water and oxygen. The Pt/Au rods were able to consistently reach speeds of up to 8 µm/s in a solution of 3.3% hydrogen peroxide. The decomposition of hydrogen peroxide in the Pt side produces oxygen, two protons and two electrons. The two protons and electrons will travel towards the Au, where they will be used to react with another hydrogen peroxide molecule, to produce two water molecules. The movements of the two protons and the two electrons through the rod drag the fluid towards the Au side, thus this fluid flow will propel the rod in the opposite direction. This self-electrophoresis mechanism is what powers the motion of these rods.[93] Further analysis of the Pt/Au rods showed that they were capable of performing chemotaxis towards higher hydrogen peroxide concentrations,[94] transport cargo,[95] and exhibited steerable motion in an external magnetic field when inner Ni segments were added.[95]
Responding to stimuli

Reconfigurable synthetic or artificial microswimmers need internal feedback[125] Self-propelling microparticles are often proposed as synthetic models for biological microswimmers, yet they lack the internally regulated adaptation of their biological counterparts. Conversely, adaptation can be encoded in larger-scale soft-robotic devices but remains elusive to transfer to the colloidal scale.[125]
The ubiquity and success of motile bacteria are strongly coupled to their ability to autonomously adapt to different environments as they can reconfigure their shape, metabolism, and motility via internal feedback mechanisms.[126][127] Realizing artificial microswimmers with similar adaptation capabilities and autonomous behavior might substantially impact technologies ranging from optimal transport to sensing and microrobotics.[128] Focusing on adaptation, existing approaches at the colloidal scale mostly rely on external feedback, either to regulate motility via the spatiotemporal modulation of the propulsion velocity and direction [129][124][130][131] or to induce shape changes via the same magnetic or electric fields,[132][133][134] which are also driving the particles. On the contrary, endowing artificial microswimmers with an internal feedback mechanism, which regulates motility in response to stimuli that are decoupled from the source of propulsion, remains an elusive task.[125]
A promising route to achieve this goal is to exploit the coupling between particle shape and motility. Efficient switching between different propulsion states can, for instance, be reached by the spontaneous aggregation of symmetry-breaking active clusters of varying geometry,[135][136][137][138] albeit this process does not have the desired deterministic control. Conversely, designing colloidal clusters with fixed shapes and compositions offers fine control on motility [139][140][141] but lacks adaptation. Although progress on reconfigurable robots at the sub-millimeter scale has been made,[142][143][144][145][146] downscaling these concepts to the colloidal level demands alternative fabrication and design. Shape-shifting colloidal clusters reconfiguring along a predefined pathway in response to local stimuli [147] would combine both characteristics, with high potential toward the vision of realising adaptive artificial microswimmers.[125]
Biohybrid microswimmers

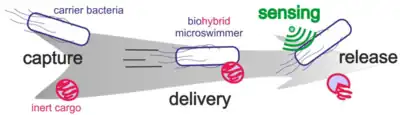
The so-called biohybrid microswimmer can be defined as a microswimmer that consist of both biological and artificial parts, for instance, one or several living microorganisms attached to one or various synthetic parts. The biohybrid approach directly employs living microorganisms to be a main component or modified base of a functional microswimmer.[150][151] Initially microorganisms were used as the motor units for artificial devices, but in recent years this role has been extended and modified toward other functionalities that take advantage of the biological capabilities of these organisms considering their means of interacting with other cells and living matter, specifically for applications inside the human body like drug delivery or fertilisation.[152][153][68]
A distinct advantage of microorganisms is that they naturally integrate motility and various biological functions in a conveniently miniaturised package, coupled with autonomous sensing and decision-making capabilities. They are able to adapt and thrive in complex in vivo environments and are capable of self-repair and self-assembly upon interaction with their surroundings. In that sense, self-sufficient microorganisms naturally function very similar to what we envision for artificially created microrobots: They harvest chemical energy from their surroundings to power molecular motor proteins that serve as actuators, they employ ion channels and microtubular networks to act as intracellular wiring, they rely on RNA or DNA as memory for control algorithms, and they feature an array of various membrane proteins to sense and evaluate their surroundings. All these abilities act together to allow microbes to thrive and pursue their goal and function. In principle, these abilities also qualify them as biological microrobots for novel operations like theranostics, the combination of diagnosis and therapy, if we are able to impose such functions artificially, for example, by functionalisation with therapeutics. Further, artificial extensions may be used as handles for external control and supervision mechanisms or to enhance the microbe's performance to guide and tailor its functions for specific applications.[68]
In fact, the biohybrid approach can be conceived in a dualistic way, with respect to the three basic ingredients of an in vivo microrobot, which are motility, control, and functionality. Figure 1 illustrates how these three ingredients can be either realized biologically, i.e., by the microorganism, or artificially, i.e., by the synthetic component. For example, a hybrid biomicromotor based on a sperm cell can be driven by the flagellum of the sperm or by an attached artificial helical flagellum.[154][155] It can orient itself autonomously via biological interactions with its surroundings and other cells, or be controlled and supervised externally via artificial sensors and actuators. Finally, it can carry out a biological function, like its inherent ability to fertilize an egg cell, or an artificially imposed function, like the delivery of synthetic drugs or DNA vectors. A biohybrid device may deploy any feasible combination of such biological and artificial components in order to carry out a specific application.[68]
Navigation
Hydrodynamics can determine the optimal route for microswimmer navigation[156] Compared to the well explored problem of how to steer a macroscopic agent, like an airplane or a moon lander, to optimally reach a target, optimal navigation strategies for microswimmers experiencing hydrodynamic interactions with walls and obstacles are far-less understood.[156] Furthermore, hydrodynamic interactions in suspensions of microswimmers produce complex behavior.[157][158] The quest on how to navigate or steer to optimally reach a target is important, e.g., for airplanes to save fuel while facing complex wind patterns on their way to a remote destination, or for the coordination of the motion of the parts of a space-agent to safely land on the moon. These classical problems are well-explored and are usually solved using optimal control theory.[159] Likewise, navigation and search strategies are frequently encountered in a plethora of biological systems, including the foraging of animals for food,[160] or of T cells searching for targets to mount an immune response.[161]
There is growing interest in optimal navigation problems and search strategies [162][163][164][165][166][167] of microswimmers [58][103][168][169] and "dry" active Brownian particles,[170][99][171][172][156] The general problem regarding the optimal trajectory of a microswimmer which can freely steer but cannot control its speed toward a predefined target (point-to-point navigation) can be referred to as "the optimal microswimmer navigation problem". The characteristic differences between the optimal microswimmer navigation problem and conventional optimal control problems for macroagents like airplanes, cruise-ships, or moon-landers root in the presence of a low-Reynolds-number solvent in the former problem only. They comprise (i) overdamped dynamics, (ii) thermal fluctuations, and (iii) long-ranged fluid-mediated hydrodynamic interactions with interfaces, walls, and obstacles, all of which are characteristic for microswimmers.[99] In particular, the non-conservative hydrodynamic forces which microswimmers experience call for a distinct navigation strategy than the conservative gravitational forces acting, e.g. on space vehicles. Recent work has explored optimal navigation problems of dry active particles (and particles in external flow fields) accounting for (i) and partly also for (ii). Specifically recent research has pioneered the use of reinforcement learning [173][174][175] such as determining optimal steering strategies of active particles to optimally navigate toward a target position [162][163][166][167] or to exploit external flow fields to avoid getting trapped in certain flow structures by learning smart gravitaxis.[176] Deep reinforcement learning has been used to explore microswimmer navigation problems in mazes and obstacle arrays [177] assuming global [163] or only local [164] knowledge of the environment. Analytical approaches to optimal active particle navigation [165][166] complement these works and allow testing machine-learned results.[166][167][156]
Applications
As is the case for microtechnology and nanotechnology in general, the history of microswimmer applications arguably starts with Richard Feynman’s famous lecture There's Plenty of Room at the Bottom.[178] In the visionary speech, among other topics, Feynman addressed the idea of microscopic surgeons, saying: "...it would be interesting in surgery if you could swallow the surgeon. You put the mechanical surgeon inside the blood vessel and it goes into the heart and <<looks>> around (of course the information has to be fed out). It finds out which valve is the faulty one and takes a little knife and slices it out. Other small machines might be permanently incorporated in the body to assist some inadequately-functioning organ." The concept of the surgeon one could swallow was soon after presented in the science-fiction movie Fantastic Voyage and in Isaac Asimov’s writings.[1]

Only a few decades later, microswimmers aiming to become true microscale surgeons evolved from an intriguing science-fiction concept to a reality explored in many research laboratories around the world, as already highlighted by Metin Sitti in 2009.[180][1] These active agents that can self-propel in a low Reynolds number environment might play a key role in the future of nanomedicine, as popularised in 2016 by Yuval Noah Harari in Homo Deus: A Brief History of Tomorrow.[181] In particular, they might become useful for the targeted delivery of genes [182] or drugs [183][184] and other cargo [185][186] to a certain target (e.g. a cancer cell) through our blood vessels, requiring them to find a good, or ideally optimal, path toward the target avoiding, e.g., obstacles and unfortunate flow field regions.[156]
Already in 2010, Nelson et al. reviewed the existing and envisioned applications of microrobots in minimally invasive medicine.[187] Since then, the field has grown, and it has become clear that microswimmers have much potential for biomedical applications.[1] Already, many interesting tasks can be performed in vitro using tailored microswimmers. Still, as of 2020, a number of challenges regarding in vivo control, biocompatibility and long-term biosafety need to be overcome before microswimmers can become a viable option for many clinical applications.[188][1]
A schematic representation of the classification of biomedical applications is shown in the diagram on the left below. This includes the use of microswimmers for cargo transport in drug delivery and other biomedical applications, as well as assisted fertilisation, sensing, micromanipulation and imaging. Some of the more complex microswimmers fit into multiple categories, as they are applied simultaneously for e.g., sensing and drug delivery.[1]


The design of an untethered microscopic mobile machine or microrobot to function in vivo with medical interventional capabilities should assume an integrated approach where design 3D body shape, material composition, manufacturing technique, deployment strategy, actuation and control methods, imaging modality, permeation of biological barriers, and the execution of the prescribed medical tasks need to be considered altogether, as illustrated in the diagram on the right above. Each of these essential aspects contains a special design consideration, which must be reflected at the physical design of the microrobot.[189]
See also
References
- 1 2 3 4 5 6 7 8 9 10 11 Bunea, Ada-Ioana; Taboryski, Rafael (2020). "Recent Advances in Microswimmers for Biomedical Applications". Micromachines. 11 (12): 1048. doi:10.3390/mi11121048. PMC 7760273. PMID 33261101.
 Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License.
Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License. - ↑ Medina-Sánchez, Mariana; Schwarz, Lukas; Meyer, Anne K.; Hebenstreit, Franziska; Schmidt, Oliver G. (2016). "Cellular Cargo Delivery: Toward Assisted Fertilization by Sperm-Carrying Micromotors". Nano Letters. 16 (1): 555–561. Bibcode:2016NanoL..16..555M. doi:10.1021/acs.nanolett.5b04221. PMID 26699202.
- 1 2 Schauer, Oliver; Mostaghaci, Babak; Colin, Remy; Hürtgen, Daniel; Kraus, David; Sitti, Metin; Sourjik, Victor (2018). "Motility and chemotaxis of bacteria-driven microswimmers fabricated using antigen 43-mediated biotin display". Scientific Reports. 8 (1): 9801. Bibcode:2018NatSR...8.9801S. doi:10.1038/s41598-018-28102-9. PMC 6023875. PMID 29955099.
- ↑ Magdanz, Veronika; Sanchez, Samuel; Schmidt, Oliver G. (2013). "Development of a Sperm-Flagella Driven Micro-Bio-Robot". Advanced Materials. 25 (45): 6581–6588. Bibcode:2013AdM....25.6581M. doi:10.1002/adma.201302544. PMID 23996782. S2CID 5125033.
- 1 2 Srivastava, Sarvesh Kumar; Medina-Sánchez, Mariana; Koch, Britta; Schmidt, Oliver G. (2016). "Medibots: Dual-Action Biogenic Microdaggers for Single-Cell Surgery and Drug Release". Advanced Materials. 28 (5): 832–837. Bibcode:2016AdM....28..832S. doi:10.1002/adma.201504327. PMID 26619085. S2CID 40955542.
- ↑ Singh, Ajay Vikram; Dad Ansari, Mohammad Hasan; Dayan, Cem Balda; Giltinan, Joshua; Wang, Shuo; Yu, Yan; Kishore, Vimal; Laux, Peter; Luch, Andreas; Sitti, Metin (2019). "Multifunctional magnetic hairbot for untethered osteogenesis, ultrasound contrast imaging and drug delivery". Biomaterials. 219: 119394. doi:10.1016/j.biomaterials.2019.119394. PMID 31382208. S2CID 199451792.
- ↑ Bhuyan, Tamanna; Singh, Amit Kumar; Dutta, Deepanjalee; Unal, Aynur; Ghosh, Siddhartha Sankar; Bandyopadhyay, Dipankar (2017). "Magnetic Field Guided Chemotaxis of i Mushbots for Targeted Anticancer Therapeutics". ACS Biomaterials Science & Engineering. 3 (8): 1627–1640. doi:10.1021/acsbiomaterials.7b00086. PMID 33429648.
- ↑ Magdanz, Veronika; Khalil, Islam S. M.; Simmchen, Juliane; Furtado, Guilherme P.; Mohanty, Sumit; Gebauer, Johannes; Xu, Haifeng; Klingner, Anke; Aziz, Azaam; Medina-Sánchez, Mariana; Schmidt, Oliver G.; Misra, Sarthak (2020). "IRONSperm: Sperm-templated soft magnetic microrobots". Science Advances. 6 (28): eaba5855. Bibcode:2020SciA....6.5855M. doi:10.1126/sciadv.aba5855. PMC 7450605. PMID 32923590.
- ↑ Bhuyan, Tamanna; Dutta, Deepanjalee; Bhattacharjee, Mitradip; Singh, Amit Kumar; Ghosh, Siddhartha Sankar; Bandyopadhyay, Dipankar (2019). "Acoustic Propulsion of Vitamin C Loaded Teabots for Targeted Oxidative Stress and Amyloid Therapeutics". ACS Applied Bio Materials. 2 (10): 4571–4582. doi:10.1021/acsabm.9b00677. PMID 35021416. S2CID 203945671.
- 1 2 3 4 Bunea, Ada-Ioana; Glückstad, Jesper (2019). "Strategies for Optical Trapping in Biological Samples: Aiming at Microrobotic Surgeons" (PDF). Laser & Photonics Reviews. 13 (4). Bibcode:2019LPRv...1300227B. doi:10.1002/lpor.201800227. S2CID 128326068.
- ↑ Bhuyan, Tamanna; Simon, Anitha T.; Maity, Surjendu; Singh, Amit Kumar; Ghosh, Siddhartha Sankar; Bandyopadhyay, Dipankar (2020). "Magnetotactic T-Budbots to Kill-n-Clean Biofilms". ACS Applied Materials & Interfaces. 12 (39): 43352–43364. doi:10.1021/acsami.0c08444. PMID 32864951. S2CID 221383266.
- ↑ Wang, Xiaopu; Chen, Xiang-Zhong; Alcântara, Carlos C. J.; Sevim, Semih; Hoop, Marcus; Terzopoulou, Anastasia; De Marco, Carmela; Hu, Chengzhi; De Mello, Andrew J.; Falcaro, Paolo; Furukawa, Shuhei; Nelson, Bradley J.; Puigmartí-Luis, Josep; Pané, Salvador (2019). "MOF-Based Microrobots: MOFBOTS: Metal–Organic-Framework-Based Biomedical Microrobots (Adv. Mater. 27/2019)". Advanced Materials. 31 (27). Bibcode:2019AdM....3170192W. doi:10.1002/adma.201970192. S2CID 198797318.
- ↑ Brown, James F. (1852). "XXIV. On some salts and products of decomposition of pyromeconic acid". The London, Edinburgh, and Dublin Philosophical Magazine and Journal of Science. 4 (24): 161–168. doi:10.1080/14786445208647098.
- ↑ Einstein, A. (1905). "Über die von der molekularkinetischen Theorie der Wärme geforderte Bewegung von in ruhenden Flüssigkeiten suspendierten Teilchen". Annalen der Physik. 322 (8): 549–560. Bibcode:1905AnP...322..549E. doi:10.1002/andp.19053220806.
- 1 2 3 4 5 6 7 8 9 10 11 12 13 14 Bastos-Arrieta, Julio; Revilla-Guarinos, Ainhoa; Uspal, William E.; Simmchen, Juliane (2018). "Bacterial Biohybrid Microswimmers". Frontiers in Robotics and AI. 5: 97. doi:10.3389/frobt.2018.00097. PMC 7805739. PMID 33500976.
 Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License.
Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License. - 1 2 3 4 5 6 7 8 Purcell, E. M. (1977). "Life at low Reynolds number". American Journal of Physics. 45 (1): 3–11. Bibcode:1977AmJPh..45....3P. doi:10.1119/1.10903.
- ↑ Qiu, Tian; Lee, Tung-Chun; Mark, Andrew G.; Morozov, Konstantin I.; Münster, Raphael; Mierka, Otto; Turek, Stefan; Leshansky, Alexander M.; Fischer, Peer (2014). "Swimming by reciprocal motion at low Reynolds number". Nature Communications. 5: 5119. Bibcode:2014NatCo...5.5119Q. doi:10.1038/ncomms6119. PMC 4241991. PMID 25369018.
- ↑ Zhang, Li; Abbott, Jake J.; Dong, Lixin; Kratochvil, Bradley E.; Bell, Dominik; Nelson, Bradley J. (2009). "Artificial bacterial flagella: Fabrication and magnetic control". Applied Physics Letters. 94 (6): 064107. Bibcode:2009ApPhL..94f4107Z. doi:10.1063/1.3079655.
- ↑ Abbott, Jake J.; Peyer, Kathrin E.; Lagomarsino, Marco Cosentino; Zhang, Li; Dong, Lixin; Kaliakatsos, Ioannis K.; Nelson, Bradley J. (2009). "How Should Microrobots Swim?". The International Journal of Robotics Research. 28 (11–12): 1434–1447. doi:10.1177/0278364909341658. S2CID 62330062.
- ↑ Schamel, Debora; Mark, Andrew G.; Gibbs, John G.; Miksch, Cornelia; Morozov, Konstantin I.; Leshansky, Alexander M.; Fischer, Peer (2014). "Nanopropellers and Their Actuation in Complex Viscoelastic Media". ACS Nano. 8 (9): 8794–8801. doi:10.1021/nn502360t. PMID 24911046.
- ↑ Rogowski, Louis William; Oxner, Micah; Tang, Jiannan; Kim, Min Jun (2020). "Heterogeneously flagellated microswimmer behavior in viscous fluids". Biomicrofluidics. 14 (2): 024112. doi:10.1063/1.5137743. PMC 7173976. PMID 32341723.
- ↑ Ceylan, Hakan; Yasa, Immihan Ceren; Yasa, Oncay; Tabak, Ahmet Fatih; Giltinan, Joshua; Sitti, Metin (2019). "3D-Printed Biodegradable Microswimmer for Theranostic Cargo Delivery and Release". ACS Nano. 13 (3): 3353–3362. doi:10.1021/acsnano.8b09233. PMC 6728090. PMID 30742410.
- ↑ Peyer, Kathrin E.; Zhang, Li; Nelson, Bradley J. (2013). "Bio-inspired magnetic swimming microrobots for biomedical applications". Nanoscale. 5 (4): 1259–1272. Bibcode:2013Nanos...5.1259P. doi:10.1039/C2NR32554C. PMID 23165991.
- ↑ Chowdhury, Sagar; Jing, Wuming; Cappelleri, David J. (2015). "Controlling multiple microrobots: Recent progress and future challenges". Journal of Micro-Bio Robotics. 10 (1–4): 1–11. doi:10.1007/s12213-015-0083-6. S2CID 53644820.
- ↑ Servant, Ania; Qiu, Famin; Mazza, Mariarosa; Kostarelos, Kostas; Nelson, Bradley J. (2015). "Controlled in Vivo Swimming of a Swarm of Bacteria-Like Microrobotic Flagella". Advanced Materials. 27 (19): 2981–2988. Bibcode:2015AdM....27.2981S. doi:10.1002/adma.201404444. PMID 25850420. S2CID 22780031.
- ↑ Dong, Xiaoguang; Sitti, Metin (2020). "Controlling two-dimensional collective formation and cooperative behavior of magnetic microrobot swarms". The International Journal of Robotics Research. 39 (5): 617–638. doi:10.1177/0278364920903107. S2CID 213942288.
- ↑ Liang, Xiong; Mou, Fangzhi; Huang, Zhen; Zhang, Jianhua; You, Ming; Xu, Leilei; Luo, Ming; Guan, Jianguo (2020). "Hierarchical Microswarms with Leader–Follower-Like Structures: Electrohydrodynamic Self-Organization and Multimode Collective Photoresponses". Advanced Functional Materials. 30 (16). doi:10.1002/adfm.201908602. S2CID 214408287.
- ↑ Zheng, Jing; Dai, Baohu; Wang, Jizhuang; Xiong, Ze; Yang, Ya; Liu, Jun; Zhan, Xiaojun; Wan, Zhihan; Tang, Jinyao (2017). "Orthogonal navigation of multiple visible-light-driven artificial microswimmers". Nature Communications. 8 (1): 1438. Bibcode:2017NatCo...8.1438Z. doi:10.1038/s41467-017-01778-9. PMC 5681650. PMID 29127414.
- 1 2 Koleoso, M.; Feng, X.; Xue, Y.; Li, Q.; Munshi, T.; Chen, X. (2020). "Micro/Nanoscale magnetic robots for biomedical applications". Materials Today Bio. 8: 100085. doi:10.1016/j.mtbio.2020.100085. PMC 7702192. PMID 33299981.
- ↑ Rao, K. Jagajjanani; Li, Fei; Meng, Long; Zheng, Hairong; Cai, Feiyan; Wang, Wei (2015). "A Force to be Reckoned with: A Review of Synthetic Microswimmers Powered by Ultrasound". Small. 11 (24): 2836–2846. doi:10.1002/smll.201403621. PMID 25851515.
- 1 2 3 Dreyfus, Rémi; Baudry, Jean; Roper, Marcus L.; Fermigier, Marc; Stone, Howard A.; Bibette, Jérôme (2005). "Microscopic artificial swimmers". Nature. 437 (7060): 862–865. Bibcode:2005Natur.437..862D. doi:10.1038/nature04090. PMID 16208366. S2CID 3025635.
- ↑ Palagi, Stefano; Mark, Andrew G.; Reigh, Shang Yik; Melde, Kai; Qiu, Tian; Zeng, Hao; Parmeggiani, Camilla; Martella, Daniele; Sanchez-Castillo, Alberto; Kapernaum, Nadia; Giesselmann, Frank; Wiersma, Diederik S.; Lauga, Eric; Fischer, Peer (2016). "Structured light enables biomimetic swimming and versatile locomotion of photoresponsive soft microrobots". Nature Materials. 15 (6): 647–653. Bibcode:2016NatMa..15..647P. doi:10.1038/nmat4569. hdl:2158/1105540. PMID 26878315.
- ↑ Kim, Min-Soo; Lee, Hyun-Taek; Ahn, Sung-Hoon (2019). "Laser Controlled 65 Micrometer Long Microrobot Made of Ni-Ti Shape Memory Alloy". Advanced Materials Technologies. 4 (12). doi:10.1002/admt.201900583. S2CID 210801365.
- ↑ Peng, Xiaolei; Chen, Zhihan; Kollipara, Pavana Siddhartha; Liu, Yaoran; Fang, Jie; Lin, Linhan; Zheng, Yuebing (2020). "Opto-thermoelectric microswimmers". Light: Science & Applications. 9 (1): 141. Bibcode:2020LSA.....9..141P. doi:10.1038/s41377-020-00378-5. PMC 7429954. PMID 32864116.
- ↑ Bastos-Arrieta, Julio; Revilla-Guarinos, Ainhoa; Uspal, William E.; Simmchen, Juliane (2018). "Bacterial Biohybrid Microswimmers". Frontiers in Robotics and AI. 5: 97. doi:10.3389/frobt.2018.00097. PMC 7805739. PMID 33500976.
- ↑ Bente, Klaas; Codutti, Agnese; Bachmann, Felix; Faivre, Damien (2018). "Biohybrid and Bioinspired Magnetic Microswimmers". Small. 14 (29): e1704374. doi:10.1002/smll.201704374. PMID 29855143. S2CID 46918320.
- ↑ Kaynak, Murat; Ozcelik, Adem; Nourhani, Amir; Lammert, Paul E.; Crespi, Vincent H.; Huang, Tony Jun (2017). "Acoustic actuation of bioinspired microswimmers". Lab on a Chip. 17 (3): 395–400. doi:10.1039/C6LC01272H. PMC 5465869. PMID 27991641.
- ↑ Xin, Hongbao; Zhao, Nan; Wang, Yunuo; Zhao, Xiaoting; Pan, Ting; Shi, Yang; Li, Baojun (2020). "Optically Controlled Living Micromotors for the Manipulation and Disruption of Biological Targets". Nano Letters. 20 (10): 7177–7185. Bibcode:2020NanoL..20.7177X. doi:10.1021/acs.nanolett.0c02501. PMID 32935992. S2CID 221747106.
- ↑ Nagai, Moeto; Hirano, Takahiro; Shibata, Takayuki (2019). "Phototactic Algae-Driven Unidirectional Transport of Submillimeter-Sized Cargo in a Microchannel". Micromachines. 10 (2): 130. doi:10.3390/mi10020130. PMC 6412834. PMID 30781488.
- ↑ Lozano, Celia; Ten Hagen, Borge; Löwen, Hartmut; Bechinger, Clemens (2016). "Phototaxis of synthetic microswimmers in optical landscapes". Nature Communications. 7: 12828. arXiv:1609.09814. Bibcode:2016NatCo...712828L. doi:10.1038/ncomms12828. PMC 5056439. PMID 27687580. S2CID 7924312.
- ↑ Singh, Dhruv P.; Uspal, William E.; Popescu, Mihail N.; Wilson, Laurence G.; Fischer, Peer (2018). "Photogravitactic Microswimmers" (PDF). Advanced Functional Materials. 28 (25). doi:10.1002/adfm.201706660. S2CID 247697846.
- ↑ Dai, Baohu; Wang, Jizhuang; Xiong, Ze; Zhan, Xiaojun; Dai, Wei; Li, Chien-Cheng; Feng, Shien-Ping; Tang, Jinyao (2016). "Programmable artificial phototactic microswimmer". Nature Nanotechnology. 11 (12): 1087–1092. Bibcode:2016NatNa..11.1087D. doi:10.1038/nnano.2016.187. PMID 27749832.
- ↑ Akolpoglu, Mukrime Birgul; Dogan, Nihal Olcay; Bozuyuk, Ugur; Ceylan, Hakan; Kizilel, Seda; Sitti, Metin (2020). "High-Yield Production of Biohybrid Microalgae for On-Demand Cargo Delivery". Advanced Science. 7 (16). doi:10.1002/advs.202001256. PMC 7435244. PMID 32832367.
- ↑ Tu, Yingfeng; Peng, Fei; Wilson, Daniela A. (2017). "Motion Manipulation of Micro- and Nanomotors". Advanced Materials. 29 (39). Bibcode:2017AdM....2901970T. doi:10.1002/adma.201701970. PMID 28841755. S2CID 205280841.
- ↑ Luo, Ming; Feng, Youzeng; Wang, Tingwei; Guan, Jianguo (2018). "Micro-/Nanorobots at Work in Active Drug Delivery". Advanced Functional Materials. 28 (25). doi:10.1002/adfm.201706100. S2CID 104145610.
- ↑ Srivastava, Sarvesh Kumar; Clergeaud, Gael; Andresen, Thomas L.; Boisen, Anja (2019). "Micromotors for drug delivery in vivo: The road ahead" (PDF). Advanced Drug Delivery Reviews. 138: 41–55. doi:10.1016/j.addr.2018.09.005. PMID 30236447. S2CID 52310451.
- ↑ Plutnar, Jan; Pumera, Martin (2019). "Chemotactic Micro- and Nanodevices". Angewandte Chemie International Edition. 58 (8): 2190–2196. doi:10.1002/anie.201809101. PMID 30216620. S2CID 52278805.
- ↑ Yang, Qingliang; Xu, Lei; Zhong, Weizhen; Yan, Qinying; Gao, Ying; Hong, Weiyong; She, Yuanbin; Yang, Gensheng (2020). "Recent Advances in Motion Control of Micro/Nanomotors". Advanced Intelligent Systems. 2 (8). doi:10.1002/aisy.202000049. S2CID 221418150.
- ↑ Kanu, Nand Jee; Gupta, Eva; Vates, Umesh Kumar; Singh, Gyanendra Kumar (2019). "An insight into biomimetic 4D printing". RSC Advances. 9 (65): 38209–38226. Bibcode:2019RSCAd...938209K. doi:10.1039/C9RA07342F. PMC 9075844. PMID 35541793. S2CID 214386444.
- ↑ Lui, Yuan Siang; Sow, Wan Ting; Tan, Lay Poh; Wu, Yunlong; Lai, Yuekun; Li, Huaqiong (2019). "4D printing and stimuli-responsive materials in biomedical aspects". Acta Biomaterialia. 92: 19–36. doi:10.1016/j.actbio.2019.05.005. hdl:10356/143207. PMID 31071476. S2CID 149445838.
- ↑ Spiegel, Christoph A.; Hippler, Marc; Münchinger, Alexander; Bastmeyer, Martin; Barner-Kowollik, Christopher; Wegener, Martin; Blasco, Eva (2020). "4D Printing at the Microscale". Advanced Functional Materials. 30 (26). doi:10.1002/adfm.201907615. S2CID 210959593.
- ↑ Yang, Qingzhen; Gao, Bin; Xu, Feng (2020). "Recent Advances in 4D Bioprinting". Biotechnology Journal. 15 (1): e1900086. doi:10.1002/biot.201900086. PMID 31486199. S2CID 201837838.
- ↑ Zhang, Yabin; Yuan, Ke; Zhang, Li (16 January 2019). "Micro/Nanomachines: from Functionalization to Sensing and Removal". Advanced Materials Technologies. Wiley. 4 (4): 1800636. doi:10.1002/admt.201800636. ISSN 2365-709X. S2CID 139612870.
- ↑ Bunea, Ada-Ioana; Jakobsen, Mogens Havsteen; Engay, Einstom; Bañas, Andrew R.; Glückstad, Jesper (2019). "Optimization of 3D-printed microstructures for investigating the properties of the mucus biobarrier". Micro and Nano Engineering. Elsevier BV. 2: 41–47. doi:10.1016/j.mne.2018.12.004. ISSN 2590-0072. S2CID 215751974.
- ↑ Zhang, Yabin; Yuan, Ke; Zhang, Li (2019). "Micro/Nanomachines: From Functionalization to Sensing and Removal". Advanced Materials Technologies. 4 (4). doi:10.1002/admt.201800636. S2CID 139612870.
- ↑ Bunea, Ada-Ioana; Jakobsen, Mogens Havsteen; Engay, Einstom; Bañas, Andrew R.; Glückstad, Jesper (2019). "Optimization of 3D-printed microstructures for investigating the properties of the mucus biobarrier". Micro and Nano Engineering. 2: 41–47. doi:10.1016/j.mne.2018.12.004. S2CID 215751974.
- ↑ Singh, Ajay Vikram; Kishore, Vimal; Santomauro, Giulia; Yasa, Oncay; Bill, Joachim; Sitti, Metin (28 April 2020). "Mechanical Coupling of Puller and Pusher Active Microswimmers Influences Motility". Langmuir. American Chemical Society (ACS). 36 (19): 5435–5443. doi:10.1021/acs.langmuir.9b03665. ISSN 0743-7463. PMC 7304893. PMID 32343587.
 Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License.
Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License. - 1 2 3 Lauga, Eric; Powers, Thomas R. (2009). "The hydrodynamics of swimming microorganisms". Reports on Progress in Physics. 72 (9): 096601. arXiv:0812.2887. Bibcode:2009RPPh...72i6601L. doi:10.1088/0034-4885/72/9/096601. S2CID 3932471.
- ↑ Vogel, Pia D. (2005). "Nature's design of nanomotors". European Journal of Pharmaceutics and Biopharmaceutics. 60 (2): 267–277. doi:10.1016/j.ejpb.2004.10.007. PMID 15939237.
- ↑ Patra, Debabrata; Sengupta, Samudra; Duan, Wentao; Zhang, Hua; Pavlick, Ryan; Sen, Ayusman (2013). "Intelligent, self-powered, drug delivery systems". Nanoscale. 5 (4): 1273–1283. Bibcode:2013Nanos...5.1273P. doi:10.1039/C2NR32600K. PMID 23166050.
- ↑ Feringa, Ben L. (2001). "In Control of Motion: From Molecular Switches to Molecular Motors". Accounts of Chemical Research. 34 (6): 504–513. doi:10.1021/ar0001721. hdl:11370/a0b20090-34b9-4e2d-8450-bc2afbea2fcf. PMID 11412087.
- ↑ Sokolov, A.; Apodaca, M. M.; Grzybowski, B. A.; Aranson, I. S. (2010). "Swimming bacteria power microscopic gears". Proceedings of the National Academy of Sciences. 107 (3): 969–974. Bibcode:2010PNAS..107..969S. doi:10.1073/pnas.0913015107. PMC 2824308. PMID 20080560.
- ↑ Zhao, Xi; Gentile, Kayla; Mohajerani, Farzad; Sen, Ayusman (2018-10-16). "Powering Motion with Enzymes". Accounts of Chemical Research. 51 (10): 2373–2381. doi:10.1021/acs.accounts.8b00286. ISSN 0001-4842. PMID 30256612. S2CID 52845451.
- ↑ Muddana, Hari S.; Sengupta, Samudra; Mallouk, Thomas E.; Sen, Ayusman; Butler, Peter J. (2010-02-24). "Substrate Catalysis Enhances Single-Enzyme Diffusion". Journal of the American Chemical Society. 132 (7): 2110–2111. doi:10.1021/ja908773a. ISSN 0002-7863. PMC 2832858. PMID 20108965.
- ↑ Ghosh, Subhadip; Mohajerani, Farzad; Son, Seoyoung; Velegol, Darrell; Butler, Peter J.; Sen, Ayusman (2019-09-11). "Motility of Enzyme-Powered Vesicles". Nano Letters. 19 (9): 6019–6026. Bibcode:2019NanoL..19.6019G. doi:10.1021/acs.nanolett.9b01830. ISSN 1530-6984. PMID 31429577.
- ↑ Madigan, Michael T.; Bender, Kelly S.; Buckley, Daniel H.; Brock, Thomas D.; Matthew Sattley, W.; Stahl, David Allan (29 January 2018). Brock Biology of Microorganisms. Pearson. ISBN 9781292235103.
- ↑ Dufrêne, Yves F. (2015). "Sticky microbes: Forces in microbial cell adhesion". Trends in Microbiology. 23 (6): 376–382. doi:10.1016/j.tim.2015.01.011. PMID 25684261.
- 1 2 3 4 5 6 Schwarz, Lukas; Medina-Sánchez, Mariana; Schmidt, Oliver G. (2017). "Hybrid Bio Micromotors". Applied Physics Reviews. 4 (3): 031301. Bibcode:2017ApPRv...4c1301S. doi:10.1063/1.4993441.
 Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License.
Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License. - ↑ Darnton, Nicholas C.; Turner, Linda; Rojevsky, Svetlana; Berg, Howard C. (2007). "On Torque and Tumbling in Swimming Escherichia coli". Journal of Bacteriology. 189 (5): 1756–1764. doi:10.1128/JB.01501-06. PMC 1855780. PMID 17189361.
- ↑ Edwards, Matthew R.; Carlsen, Rika Wright; Zhuang, Jiang; Sitti, Metin (2014). "Swimming characterization of Serratia marcescens for bio-hybrid micro-robotics". Journal of Micro-Bio Robotics. 9 (3–4): 47–60. doi:10.1007/s12213-014-0072-1. S2CID 84413776.
- ↑ Magariyama, Yukio; Sugiyama, Shigeru; Kudo, Seishi (2001). "Bacterial swimming speed and rotation rate of bundled flagella". FEMS Microbiology Letters. 199 (1): 125–129. doi:10.1111/j.1574-6968.2001.tb10662.x. PMID 11356579.
- ↑ Ito, Masahiro; Terahara, Naoya; Fujinami, Shun; Krulwich, Terry Ann (2005). "Properties of Motility in Bacillus subtilis Powered by the H+-coupled MotAB Flagellar Stator, Na+-coupled MotPS or Hybrid Stators MotAS or MotPB". Journal of Molecular Biology. 352 (2): 396–408. doi:10.1016/j.jmb.2005.07.030. PMC 2578835. PMID 16095621.
- ↑ Higashi, Kazuhiko; Miki, Norihisa (2014). "A self-swimming microbial robot using microfabricated nanofibrous hydrogel". Sensors and Actuators B: Chemical. 202: 301–306. doi:10.1016/j.snb.2014.05.068.
- ↑ Kawagishi, I.; Maekawa, Y.; Atsumi, T.; Homma, M.; Imae, Y. (1995). "Isolation of the polar and lateral flagellum-defective mutants in Vibrio alginolyticus and identification of their flagellar driving energy sources". Journal of Bacteriology. 177 (17): 5158–5160. doi:10.1128/jb.177.17.5158-5160.1995. PMC 177299. PMID 7665498.
- ↑ Xie, L.; Altindal, T.; Chattopadhyay, S.; Wu, X.-L. (2011). "Bacterial flagellum as a propeller and as a rudder for efficient chemotaxis". Proceedings of the National Academy of Sciences. 108 (6): 2246–2251. doi:10.1073/pnas.1011953108. PMC 3038696. PMID 21205908.
- ↑ Lacayo, Catherine I.; Theriot, Julie A. (2004). "Listeria monocytogenes Actin-based Motility Varies Depending on Subcellular Location: A Kinematic Probe for Cytoarchitecture". Molecular Biology of the Cell. 15 (5): 2164–2175. doi:10.1091/mbc.E03-10-0747. PMC 404013. PMID 15004231.
- ↑ McGrath, James L.; Eungdamrong, Narat J.; Fisher, Charles I.; Peng, Fay; Mahadevan, Lakshminarayanan; Mitchison, Timothy J.; Kuo, Scot C. (2003). "The Force-Velocity Relationship for the Actin-Based Motility of Listeria monocytogenes". Current Biology. 13 (4): 329–332. doi:10.1016/S0960-9822(03)00051-4. PMID 12593799. S2CID 6459972.
- ↑ Chen, Yifan; Kosmas, Panagiotis; Martel, Sylvain (2013). "A Feasibility Study for Microwave Breast Cancer Detection Using Contrast-Agent-Loaded Bacterial Microbots". International Journal of Antennas and Propagation. 2013: 1–11. doi:10.1155/2013/309703.
- ↑ Ruan, J.; Kato, T.; Santini, C.-L.; Miyata, T.; Kawamoto, A.; Zhang, W.-J.; Bernadac, A.; Wu, L.-F.; Namba, K. (2012). "Architecture of a flagellar apparatus in the fast-swimming magnetotactic bacterium MO-1". Proceedings of the National Academy of Sciences. 109 (50): 20643–20648. Bibcode:2012PNAS..10920643R. doi:10.1073/pnas.1215274109. PMC 3528567. PMID 23184985.
- ↑ Martel, Sylvain; Tremblay, Charles C.; Ngakeng, Serge; Langlois, Guillaume (2006). "Controlled manipulation and actuation of micro-objects with magnetotactic bacteria". Applied Physics Letters. 89 (23): 233904. Bibcode:2006ApPhL..89w3904M. doi:10.1063/1.2402221.
- ↑ Miyata, Makoto; Ryu, William S.; Berg, Howard C. (2002). "Force and Velocity of Mycoplasma mobile Gliding". Journal of Bacteriology. 184 (7): 1827–1831. doi:10.1128/JB.184.7.1827-1831.2002. PMC 134919. PMID 11889087.
- ↑ Weibel, D. B.; Garstecki, P.; Ryan, D.; Diluzio, W. R.; Mayer, M.; Seto, J. E.; Whitesides, G. M. (2005). "Microoxen: Microorganisms to move microscale loads". Proceedings of the National Academy of Sciences. 102 (34): 11963–11967. Bibcode:2005PNAS..10211963W. doi:10.1073/pnas.0505481102. PMC 1189341. PMID 16103369.
- ↑ Kim, Dal Hyung; Cheang, U. Kei; Kőhidai, László; Byun, Doyoung; Kim, Min Jun (2010). "Artificial magnetotactic motion control of Tetrahymena pyriformis using ferromagnetic nanoparticles: A tool for fabrication of microbiorobots". Applied Physics Letters. 97 (17): 173702. Bibcode:2010ApPhL..97q3702K. doi:10.1063/1.3497275.
- ↑ Hill, Kent L. (2003). "Biology and Mechanism of Trypanosome Cell Motility". Eukaryotic Cell. 2 (2): 200–208. doi:10.1128/EC.2.2.200-208.2003. PMC 154846. PMID 12684369.
- ↑ Krüger, Timothy; Engstler, Markus (2016). "Trypanosomes – versatile microswimmers". The European Physical Journal Special Topics. 225 (11–12): 2157–2172. Bibcode:2016EPJST.225.2157K. doi:10.1140/epjst/e2016-60063-5. S2CID 125623927.
- 1 2 Maree, L.; Van Der Horst, G. (2013). "Quantification and identification of sperm subpopulations using computer-aided sperm analysis and species-specific cut-off values for swimming speed". Biotechnic & Histochemistry. 88 (3–4): 181–193. doi:10.3109/10520295.2012.757366. hdl:10566/3120. PMID 23331185. S2CID 19603301.
- 1 2 Eamer, Lise; Nosrati, Reza; Vollmer, Marion; Zini, Armand; Sinton, David (2015). "Microfluidic assessment of swimming media for motility-based sperm selection". Biomicrofluidics. 9 (4): 044113. doi:10.1063/1.4928129. PMC 4529441. PMID 26339314.
- 1 2 Gomendio, Montserrat; Roldan, Eduardo R.S. (2008). "Implications of diversity in sperm size and function for sperm competition and fertility". The International Journal of Developmental Biology. 52 (5–6): 439–447. doi:10.1387/ijdb.082595mg. PMID 18649256.
- ↑ Tung, Chih-Kuan; Ardon, Florencia; Fiore, Alyssa G.; Suarez, Susan S.; Wu, Mingming (2014). "Cooperative roles of biological flow and surface topography in guiding sperm migration revealed by a microfluidic model". Lab Chip. 14 (7): 1348–1356. doi:10.1039/C3LC51297E. PMC 4497544. PMID 24535032.
- ↑ Ishikawa, Takuji (2019) Special Issue "Microswimmer" Micromachines, ISSN 2072-666X.
- ↑ Dumé, Isabelle (2020) Microswimmers benefit from thermoelectric guidance Physics World.
- 1 2 Paxton, Walter F.; Kistler, Kevin C.; Olmeda, Christine C.; Sen, Ayusman; St. Angelo, Sarah K.; Cao, Yanyan; Mallouk, Thomas E.; Lammert, Paul E.; Crespi, Vincent H. (2004-10-01). "Catalytic Nanomotors: Autonomous Movement of Striped Nanorods". Journal of the American Chemical Society. 126 (41): 13424–13431. doi:10.1021/ja047697z. ISSN 0002-7863. PMID 15479099.
- 1 2 Paxton, Walter F.; Baker, Paul T.; Kline, Timothy R.; Wang, Yang; Mallouk, Thomas E.; Sen, Ayusman (2006-11-01). "Catalytically Induced Electrokinetics for Motors and Micropumps". Journal of the American Chemical Society. 128 (46): 14881–14888. doi:10.1021/ja0643164. ISSN 0002-7863. PMID 17105298.
- 1 2 Hong, Yiying; Blackman, Nicole M. K.; Kopp, Nathaniel D.; Sen, Ayusman; Velegol, Darrell (2007-10-26). "Chemotaxis of Nonbiological Colloidal Rods". Physical Review Letters. 99 (17): 178103. Bibcode:2007PhRvL..99q8103H. doi:10.1103/PhysRevLett.99.178103. PMID 17995374.
- 1 2 3 Sundararajan, Shakuntala; Lammert, Paul E.; Zudans, Andrew W.; Crespi, Vincent H.; Sen, Ayusman (2008-05-01). "Catalytic Motors for Transport of Colloidal Cargo". Nano Letters. 8 (5): 1271–1276. Bibcode:2008NanoL...8.1271S. doi:10.1021/nl072275j. ISSN 1530-6984. PMID 18416540.
- ↑ Zhou, Dekai; Ren, Liqiang; Li, Yuguang C.; Xu, Pengtao; Gao, Yuan; Zhang, Guangyu; Wang, Wei; Mallouk, Thomas E.; Li, Longqiu (2017). "Visible light-driven, magnetically steerable gold/iron oxide nanomotors". Chem. Commun. 53 (83): 11465–11468. doi:10.1039/C7CC06327J. ISSN 1359-7345. PMID 28983536.
- ↑ Wang, Wei; Castro, Luz Angelica; Hoyos, Mauricio; Mallouk, Thomas (2012). "Autonomous motion of metallic microrods propelled by ultrasound". ACS Nano. 6 (7): 6122–6132. doi:10.1021/nn301312z. PMID 22631222.
- ↑ Guix, Maria; Mayorga-Martinez, Carmen C.; Merkoçi, Arben (2014). "Nano/Micromotors in (Bio)chemical Science Applications". Chemical Reviews. 114 (12): 6285–6322. doi:10.1021/cr400273r. PMID 24827167.
- 1 2 3 4 Bechinger, Clemens; Di Leonardo, Roberto; Löwen, Hartmut; Reichhardt, Charles; Volpe, Giorgio; Volpe, Giovanni (2016). "Active Particles in Complex and Crowded Environments". Reviews of Modern Physics. 88 (4): 045006. arXiv:1602.00081. Bibcode:2016RvMP...88d5006B. doi:10.1103/RevModPhys.88.045006. S2CID 14940249.
- ↑ Magdanz, Veronika; Guix, Maria; Schmidt, Oliver G. (2014). "Tubular micromotors: From microjets to spermbots". Robotics and Biomimetics. 1. doi:10.1186/s40638-014-0011-6. S2CID 55870000.
- ↑ McNeill, Jeffrey M.; Mallouk, Thomas E. (2023-10-14). "Acoustically Powered Nano- and Microswimmers: From Individual to Collective Behavior". ACS Nanoscience Au. 3 (6): 424–440. doi:10.1021/acsnanoscienceau.3c00038. ISSN 2694-2496. PMC 10740144. PMID 38144701.
- ↑ Ricotti, Leonardo; Cafarelli, Andrea; Iacovacci, Veronica; Vannozzi, Lorenzo; Menciassi, Arianna (2015). "Advanced Micro-Nano-Bio Systems for Future Targeted Therapies". Current Nanoscience. 11 (2): 144–160. Bibcode:2015CNan...11..144R. doi:10.2174/1573413710666141114221246.
- 1 2 Elgeti, J.; Winkler, R. G.; Gompper, G. (2015). "Physics of microswimmers—single particle motion and collective behavior: A review". Reports on Progress in Physics. 78 (5): 056601. arXiv:1412.2692. Bibcode:2015RPPh...78e6601E. doi:10.1088/0034-4885/78/5/056601. PMID 25919479. S2CID 3909877.
- 1 2 Purcell, E. M. (1997). "The efficiency of propulsion by a rotating flagellum". Proceedings of the National Academy of Sciences. 94 (21): 11307–11311. Bibcode:1997PNAS...9411307P. doi:10.1073/pnas.94.21.11307. PMC 23452. PMID 9326605.
- ↑ Morozov, Konstantin I.; Leshansky, Alexander M. (2014). "The chiral magnetic nanomotors". Nanoscale. 6 (3): 1580–1588. arXiv:1308.6115. Bibcode:2014Nanos...6.1580M. doi:10.1039/C3NR04853E. PMID 24336860. S2CID 15834620.
- 1 2 Lauga, Eric; Powers, Thomas R (25 August 2009). "The hydrodynamics of swimming microorganisms". Reports on Progress in Physics. IOP Publishing. 72 (9): 096601. arXiv:0812.2887. Bibcode:2009RPPh...72i6601L. doi:10.1088/0034-4885/72/9/096601. ISSN 0034-4885. S2CID 3932471.
- 1 2 Berg, Howard C.; Anderson, Robert A. (1973). "Bacteria Swim by Rotating their Flagellar Filaments". Nature. 245 (5425): 380–382. Bibcode:1973Natur.245..380B. doi:10.1038/245380a0. PMID 4593496. S2CID 4173914.
- ↑ Berg, Howard (2004). E. coli in motion (in Italian). New York: Springer. ISBN 978-0-387-21638-6. OCLC 56124142.
- ↑ Mitchell, David R. (2001). "Chlamydomonas flagella". Journal of Phycology. 36 (2): 261–273. doi:10.1046/j.1529-8817.2000.99218.x. S2CID 221921243.
- ↑ Oberholzer, Michael; Lopez, Miguel A.; McLelland, Bryce T.; Hill, Kent L. (2010). "Social Motility in African Trypanosomes". PLOS Pathogens. 6 (1): e1000739. doi:10.1371/journal.ppat.1000739. PMC 2813273. PMID 20126443.
- ↑ Babu, Sujin B.; Stark, Holger (2012). "Modeling the locomotion of the African trypanosome using multi-particle collision dynamics". New Journal of Physics. 14 (8): 085012. Bibcode:2012NJPh...14h5012B. doi:10.1088/1367-2630/14/8/085012.
- 1 2 3 Choudhary, Priyanka; Mandal, Subhayan; Babu, Sujin B. (2018). "Locomotion of a flexible one-hinge swimmer in Stokes regime". Journal of Physics Communications. 2 (2): 025009. arXiv:1707.07451. Bibcode:2018JPhCo...2b5009C. doi:10.1088/2399-6528/aaa856. S2CID 119229534.
 Material was copied from this source, which is available under a Creative Commons Attribution 3.0 International License.
Material was copied from this source, which is available under a Creative Commons Attribution 3.0 International License. - ↑ Taylor, Geoffrey (1951). "Analysis of the swimming of microscopic organisms". Proceedings of the Royal Society of London. Series A. Mathematical and Physical Sciences. 209 (1099): 447–461. Bibcode:1951RSPSA.209..447T. doi:10.1098/rspa.1951.0218. S2CID 120382159.
- ↑ Avron, J. E.; Raz, O. (2008). "A geometric theory of swimming: Purcell's swimmer and its symmetrized cousin". New Journal of Physics. 10 (6): 063016. arXiv:0712.2047. Bibcode:2008NJPh...10f3016A. doi:10.1088/1367-2630/10/6/063016. S2CID 14646885.
- ↑ Najafi, Ali; Golestanian, Ramin (2004). "Simple swimmer at low Reynolds number: Three linked spheres". Physical Review E. 69 (6): 062901. arXiv:cond-mat/0402070. Bibcode:2004PhRvE..69f2901N. doi:10.1103/PhysRevE.69.062901. PMID 15244646. S2CID 27500334.
- ↑ Daddi-Moussa-Ider, Abdallah; Lisicki, Maciej; Mathijssen, Arnold J. T. M. (2020). "Tuning the upstream swimming of microrobots by shape and cargo size". Physical Review Applied. 14 (2): 024071. arXiv:2004.05694. Bibcode:2020PhRvP..14b4071D. doi:10.1103/PhysRevApplied.14.024071. S2CID 229547570.
- ↑ Daddi-Moussa-Ider, Abdallah; Lisicki, Maciej; Hoell, Christian; Löwen, Hartmut (2018). "Swimming trajectories of a three-sphere microswimmer near a wall". Journal of Chemical Physics. 148 (13): 134904. arXiv:1801.01162. Bibcode:2018JChPh.148m4904D. doi:10.1063/1.5021027. PMID 29626882. S2CID 4718416.
- ↑ Nasouri, Babak; Khot, Aditi; Elfring, Gwynn J. (2017). "Elastic two-sphere swimmer in Stokes flow". Physical Review Fluids. 2 (4): 043101. arXiv:1611.05847. Bibcode:2017PhRvF...2d3101N. doi:10.1103/PhysRevFluids.2.043101. S2CID 119474335.
- ↑ Montino, Alessandro; Desimone, Antonio (2015). "Three-sphere low-Reynolds-number swimmer with a passive elastic arm". The European Physical Journal E. 38 (5): 127. doi:10.1140/epje/i2015-15042-3. PMID 25990633. S2CID 45431975.
- ↑ Wiggins, Chris H.; Goldstein, Raymond E. (1998). "Flexive and Propulsive Dynamics of Elastica at Low Reynolds Number". Physical Review Letters. 80 (17): 3879–3882. arXiv:cond-mat/9707346. Bibcode:1998PhRvL..80.3879W. doi:10.1103/PhysRevLett.80.3879. S2CID 10335181.
- ↑ Lagomarsino, M.C.; Capuani, F.; Lowe, C.P. (2003). "A simulation study of the dynamics of a driven filament in an Aristotelian fluid". Journal of Theoretical Biology. 224 (2): 215–224. Bibcode:2003JThBi.224..215L. doi:10.1016/S0022-5193(03)00159-0. hdl:2434/802791. PMID 12927528. S2CID 3200289.
- ↑ Lauga, Eric (2007). "Floppy swimming: Viscous locomotion of actuated elastica". Physical Review E. 75 (4): 041916. arXiv:cond-mat/0610154. Bibcode:2007PhRvE..75d1916L. doi:10.1103/PhysRevE.75.041916. PMID 17500930. S2CID 13651250.
- ↑ Ismagilov, Rustem F.; Schwartz, Alexander; Bowden, Ned; Whitesides, George M. (2002-02-15). "Autonomous Movement and Self-Assembly". Angewandte Chemie International Edition. 41 (4): 652–654. doi:10.1002/1521-3773(20020215)41:4<652::AID-ANIE652>3.0.CO;2-U. ISSN 1433-7851.
- 1 2 Khadka, Utsab; Holubec, Viktor; Yang, Haw; Cichos, Frank (2018). "Active particles bound by information flows". Nature Communications. 9 (1): 3864. arXiv:1803.03053. Bibcode:2018NatCo...9.3864K. doi:10.1038/s41467-018-06445-1. PMC 6154969. PMID 30242284.
 Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License.
Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License. - 1 2 3 4 Alvarez, L.; Fernandez-Rodriguez, M. A.; Alegria, A.; Arrese-Igor, S.; Zhao, K.; Kröger, M.; Isa, Lucio (2021). "Reconfigurable artificial microswimmers with internal feedback". Nature Communications. 12 (1): 4762. arXiv:2009.08382. Bibcode:2021NatCo..12.4762A. doi:10.1038/s41467-021-25108-2. PMC 8346629. PMID 34362934.
 Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License.
Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License. - ↑ Hamadeh, Abdullah; Roberts, Mark A. J.; August, Elias; McSharry, Patrick E.; Maini, Philip K.; Armitage, Judith P.; Papachristodoulou, Antonis (2011). "Feedback Control Architecture and the Bacterial Chemotaxis Network". PLOS Computational Biology. 7 (5): e1001130. Bibcode:2011PLSCB...7E1130H. doi:10.1371/journal.pcbi.1001130. PMC 3088647. PMID 21573199.
- ↑ Baker, Melinda D.; Wolanin, Peter M.; Stock, Jeffry B. (2006). "Signal transduction in bacterial chemotaxis". BioEssays. 28 (1): 9–22. doi:10.1002/bies.20343. PMID 16369945. S2CID 189870.
- ↑ Ebbens, S.J. (2016). "Active colloids: Progress and challenges towards realising autonomous applications". Current Opinion in Colloid & Interface Science. 21: 14–23. doi:10.1016/j.cocis.2015.10.003.
- ↑ Lozano, Celia; Ten Hagen, Borge; Löwen, Hartmut; Bechinger, Clemens (2016). "Phototaxis of synthetic microswimmers in optical landscapes". Nature Communications. 7: 12828. arXiv:1609.09814. Bibcode:2016NatCo...712828L. doi:10.1038/ncomms12828. PMC 5056439. PMID 27687580.
- ↑ Sprenger, Alexander R.; Fernandez-Rodriguez, Miguel Angel; Alvarez, Laura; Isa, Lucio; Wittkowski, Raphael; Löwen, Hartmut (2020). "Active Brownian Motion with Orientation-Dependent Motility: Theory and Experiments". Langmuir. 36 (25): 7066–7073. arXiv:1911.09524. doi:10.1021/acs.langmuir.9b03617. PMID 31975603. S2CID 208201932.
- ↑ Fernandez-Rodriguez, Miguel Angel; Grillo, Fabio; Alvarez, Laura; Rathlef, Marco; Buttinoni, Ivo; Volpe, Giovanni; Isa, Lucio (2020). "Feedback-controlled active brownian colloids with space-dependent rotational dynamics". Nature Communications. 11 (1): 4223. arXiv:1911.02291. Bibcode:2020NatCo..11.4223F. doi:10.1038/s41467-020-17864-4. PMC 7445303. PMID 32839447.
- ↑ Han, Koohee; Shields, C. Wyatt; Diwakar, Nidhi M.; Bharti, Bhuvnesh; López, Gabriel P.; Velev, Orlin D. (2017). "Sequence-encoded colloidal origami and microbot assemblies from patchy magnetic cubes". Science Advances. 3 (8): e1701108. Bibcode:2017SciA....3E1108H. doi:10.1126/sciadv.1701108. PMC 5544397. PMID 28798960.
- ↑ Shields, C. Wyatt; Velev, Orlin D. (2017). "The Evolution of Active Particles: Toward Externally Powered Self-Propelling and Self-Reconfiguring Particle Systems". Chem. 3 (4): 539–559. doi:10.1016/j.chempr.2017.09.006.
- ↑ Yang, Tao; Sprinkle, Brennan; Guo, Yang; Qian, Jun; Hua, Daoben; Donev, Aleksandar; Marr, David W. M.; Wu, Ning (2020). "Reconfigurable microbots folded from simple colloidal chains". Proceedings of the National Academy of Sciences. 117 (31): 18186–18193. Bibcode:2020PNAS..11718186Y. doi:10.1073/pnas.2007255117. PMC 7414297. PMID 32680965.
- ↑ Soto, Rodrigo; Golestanian, Ramin (2014). "Self-Assembly of Catalytically Active Colloidal Molecules: Tailoring Activity Through Surface Chemistry". Physical Review Letters. 112 (6): 068301. arXiv:1306.6596. Bibcode:2014PhRvL.112f8301S. doi:10.1103/PhysRevLett.112.068301. PMID 24580712. S2CID 37057964.
- ↑ Niu, Ran; Fischer, Andreas; Palberg, Thomas; Speck, Thomas (2018). "Dynamics of Binary Active Clusters Driven by Ion-Exchange Particles". ACS Nano. 12 (11): 10932–10938. doi:10.1021/acsnano.8b04221. PMID 30346687. S2CID 206722021.
- ↑ Ma, Fuduo; Wang, Sijia; Wu, David T.; Wu, Ning (2015). "Electric-field–induced assembly and propulsion of chiral colloidal clusters". Proceedings of the National Academy of Sciences. 112 (20): 6307–6312. Bibcode:2015PNAS..112.6307M. doi:10.1073/pnas.1502141112. PMC 4443365. PMID 25941383.
- ↑ Wang, Zuochen; Wang, Zhisheng; Li, Jiahui; Tian, Changhao; Wang, Yufeng (2020). "Active colloidal molecules assembled via selective and directional bonds". Nature Communications. 11 (1): 2670. Bibcode:2020NatCo..11.2670W. doi:10.1038/s41467-020-16506-z. PMC 7260206. PMID 32471993.
- ↑ Ebbens, Stephen; Jones, Richard A. L.; Ryan, Anthony J.; Golestanian, Ramin; Howse, Jonathan R. (2010). "Self-assembled autonomous runners and tumblers". Physical Review E. 82 (1 Pt 2): 015304. Bibcode:2010PhRvE..82a5304E. doi:10.1103/PhysRevE.82.015304. PMID 20866681.
- ↑ Ni, Songbo; Marini, Emanuele; Buttinoni, Ivo; Wolf, Heiko; Isa, Lucio (2017). "Hybrid colloidal microswimmers through sequential capillary assembly". Soft Matter. 13 (23): 4252–4259. doi:10.1039/c7sm00443e. PMID 28573270.
- ↑ Wang, Zuochen; Wang, Zhisheng; Li, Jiahui; Cheung, Simon Tsz Hang; Tian, Changhao; Kim, Shin-Hyun; Yi, Gi-Ra; Ducrot, Etienne; Wang, Yufeng (2019). "Active Patchy Colloids with Shape-Tunable Dynamics". Journal of the American Chemical Society. 141 (37): 14853–14863. doi:10.1021/jacs.9b07785. PMID 31448592. S2CID 201748635.
- ↑ Hu, Chengzhi; Pané, Salvador; Nelson, Bradley J. (2018). "Soft Micro- and Nanorobotics". Annual Review of Control, Robotics, and Autonomous Systems. 1: 53–75. doi:10.1146/annurev-control-060117-104947. hdl:20.500.11850/316345. S2CID 139844553.
- ↑ Palagi, Stefano; Fischer, Peer (2018). "Bioinspired microrobots". Nature Reviews Materials. 3 (6): 113–124. Bibcode:2018NatRM...3..113P. doi:10.1038/s41578-018-0016-9. S2CID 189929035.
- ↑ Medina-Sánchez, Mariana; Magdanz, Veronika; Guix, Maria; Fomin, Vladimir M.; Schmidt, Oliver G. (2018). "Swimming Microrobots: Soft, Reconfigurable, and Smart". Advanced Functional Materials. 28 (25). doi:10.1002/adfm.201707228. S2CID 103866599.
- ↑ Hu, Wenqi; Lum, Guo Zhan; Mastrangeli, Massimo; Sitti, Metin (2018). "Small-scale soft-bodied robot with multimodal locomotion". Nature. 554 (7690): 81–85. Bibcode:2018Natur.554...81H. doi:10.1038/nature25443. PMID 29364873. S2CID 4461200.
- ↑ Huang, H.-W.; Uslu, F. E.; Katsamba, P.; Lauga, E.; Sakar, M. S.; Nelson, B. J.; Nelson, Bradley J. (2019). "Adaptive locomotion of artificial microswimmers". Science Advances. 5 (1): eaau1532. arXiv:1902.09000. Bibcode:2019SciA....5.1532H. doi:10.1126/sciadv.aau1532. PMC 6357760. PMID 30746446.
- ↑ Dou, Yong; Bishop, Kyle J. M. (2019). "Autonomous navigation of shape-shifting microswimmers". Physical Review Research. 1 (3): 032030. arXiv:1908.05808. Bibcode:2019PhRvR...1c2030D. doi:10.1103/PhysRevResearch.1.032030. S2CID 201058417.
- ↑ Zhuang, Jiang; Park, Byung-Wook; Sitti, Metin (2017). "Propulsion and Chemotaxis in Bacteria-Driven Microswimmers". Advanced Science. 4 (9). doi:10.1002/advs.201700109. PMC 5604384. PMID 28932674.
 Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License.
Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License. - ↑ Sun, Zhiyong; Popp, Philipp; Loderer, Christoph; Revilla-Guarinos, Ainhoa (28 December 2019). "Genetically Engineered Bacterial Biohybrid Microswimmers for Sensing Applications". Sensors. MDPI AG. 20 (1): 180. Bibcode:2019Senso..20..180S. doi:10.3390/s20010180. ISSN 1424-8220. PMC 6982730. PMID 31905650.
 Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License
Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License - ↑ Carlsen, Rika Wright; Sitti, Metin (2014). "Bio-Hybrid Cell-Based Actuators for Microsystems". Small. 10 (19): 3831–3851. doi:10.1002/smll.201400384. PMID 24895215.
- ↑ Hosseinidoust, Zeinab; Mostaghaci, Babak; Yasa, Oncay; Park, Byung-Wook; Singh, Ajay Vikram; Sitti, Metin (2016). "Bioengineered and biohybrid bacteria-based systems for drug delivery". Advanced Drug Delivery Reviews. 106 (Pt A): 27–44. doi:10.1016/j.addr.2016.09.007. PMID 27641944.
- ↑ Magdanz, Veronika; Medina-Sánchez, Mariana; Schwarz, Lukas; Xu, Haifeng; Elgeti, Jens; Schmidt, Oliver G. (2017). "Spermatozoa as Functional Components of Robotic Microswimmers". Advanced Materials. 29 (24). Bibcode:2017AdM....2906301M. doi:10.1002/adma.201606301. PMID 28323360. S2CID 26622101.
- ↑ Medina-Sánchez, Mariana; Schmidt, Oliver G. (2017). "Medical microbots need better imaging and control". Nature. 545 (7655): 406–408. Bibcode:2017Natur.545..406M. doi:10.1038/545406a. PMID 28541344. S2CID 4388403.
- ↑ Magdanz, Veronika; Medina-Sánchez, Mariana; Schwarz, Lukas; Xu, Haifeng; Elgeti, Jens; Schmidt, Oliver G. (2017). "Spermatozoa as Functional Components of Robotic Microswimmers". Advanced Materials. 29 (24). Bibcode:2017AdM....2906301M. doi:10.1002/adma.201606301. PMID 28323360. S2CID 26622101.
- ↑ Medina-Sánchez, Mariana; Schmidt, Oliver G. (2017). "Medical microbots need better imaging and control". Nature. 545 (7655): 406–408. Bibcode:2017Natur.545..406M. doi:10.1038/545406a. PMID 28541344. S2CID 4388403.
- 1 2 3 4 5 Daddi-Moussa-Ider, Abdallah; Löwen, Hartmut; Liebchen, Benno (2021). "Hydrodynamics can determine the optimal route for microswimmer navigation". Communications Physics. 4 (1): 15. arXiv:2008.11064. Bibcode:2021CmPhy...4...15D. doi:10.1038/s42005-021-00522-6. S2CID 234012727.
 Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License.
Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License. - ↑ Schwarzendahl, Fabian Jan; Mazza, Marco G. (2018). "Maximum in density heterogeneities of active swimmers". Soft Matter. 14 (23): 4666–4678. arXiv:1711.08689. Bibcode:2018SMat...14.4666S. doi:10.1039/C7SM02301D. PMID 29717736.
- ↑ Theers, Mario; Westphal, Elmar; Qi, Kai; Winkler, Roland G.; Gompper, Gerhard (31 October 2018). "Clustering of microswimmers: interplay of shape and hydrodynamics". Soft Matter. 14 (42): 8590–8603. arXiv:1807.01211. Bibcode:2018SMat...14.8590T. doi:10.1039/C8SM01390J. PMID 30339172.
- ↑ Kirk, Donald (2004). Optimal control theory : an introduction. Mineola, N.Y. ISBN 978-0-486-13507-6.
{{cite book}}: CS1 maint: location missing publisher (link) - ↑ Viswanathan, Gandhimohan. M.; Da Luz, Marcos G. E.; Raposo, Ernesto P.; Stanley, H. Eugene (2011). The Physics of Foraging. doi:10.1017/CBO9780511902680. ISBN 9780511902680.
- ↑ Fricke, G. Matthew; Letendre, Kenneth A.; Moses, Melanie E.; Cannon, Judy L. (2016). "Persistence and Adaptation in Immunity: T Cells Balance the Extent and Thoroughness of Search". PLOS Computational Biology. 12 (3): e1004818. Bibcode:2016PLSCB..12E4818F. doi:10.1371/journal.pcbi.1004818. PMC 4798282. PMID 26990103.
- 1 2 Muiños-Landin, S.; Fischer, A.; Holubec, V.; Cichos, F. (2021). "Reinforcement learning with artificial microswimmers". Science Robotics. 6 (52). arXiv:1803.06425. doi:10.1126/scirobotics.abd9285. PMID 34043550. S2CID 4938282.
- 1 2 3 Yang, Yuguang; Bevan, Michael A. (2018). "Optimal Navigation of Self-Propelled Colloids". ACS Nano. 12 (11): 10712–10724. doi:10.1021/acsnano.8b05371. PMID 30252442. S2CID 52824752.
- 1 2 Yang, Yuguang; Bevan, Michael A.; Li, Bo (2020). "Efficient Navigation of Colloidal Robots in an Unknown Environment via Deep Reinforcement Learning". Advanced Intelligent Systems. 2. arXiv:1906.10844. doi:10.1002/aisy.201900106. S2CID 199000857.
- 1 2 Liebchen, Benno; Löwen, Hartmut (2019). "Optimal navigation strategies for active particles". EPL (Europhysics Letters). 127 (3): 34003. Bibcode:2019EL....12734003L. doi:10.1209/0295-5075/127/34003. S2CID 203038971.
- 1 2 3 4 Schneider, E.; Stark, H. (2019). "Optimal steering of a smart active particle". EPL (Europhysics Letters). 127 (6): 64003. arXiv:1909.03243. Bibcode:2019EL....12764003S. doi:10.1209/0295-5075/127/64003. S2CID 202540395.
- 1 2 3 Biferale, L.; Bonaccorso, F.; Buzzicotti, M.; Clark Di Leoni, P.; Gustavsson, K. (2019). "Zermelo's problem: Optimal point-to-point navigation in 2D turbulent flows using reinforcement learning". Chaos: An Interdisciplinary Journal of Nonlinear Science. 29 (10): 103138. arXiv:1907.08591. Bibcode:2019Chaos..29j3138B. doi:10.1063/1.5120370. PMID 31675828. S2CID 197935446.
- ↑ Lauga, Eric (2016). "Bacterial Hydrodynamics". Annual Review of Fluid Mechanics. 48 (1): 105–130. arXiv:1509.02184. Bibcode:2016AnRFM..48..105L. doi:10.1146/annurev-fluid-122414-034606. S2CID 13849152.
- ↑ Lauga, Eric (2020). The fluid dynamics of cell motility. Cambridge, United Kingdom New York, NY. ISBN 978-1-107-17465-8.
{{cite book}}: CS1 maint: location missing publisher (link) - ↑ Romanczuk, P.; Bär, M.; Ebeling, W.; Lindner, B.; Schimansky-Geier, L. (2012). "Active Brownian particles". The European Physical Journal Special Topics. 202: 1–162. arXiv:1202.2442. doi:10.1140/epjst/e2012-01529-y. S2CID 119100040.
- ↑ Cates, Michael E.; Tailleur, Julien (2015). "Motility-Induced Phase Separation". Annual Review of Condensed Matter Physics. 6: 219–244. arXiv:1406.3533. Bibcode:2015ARCMP...6..219C. doi:10.1146/annurev-conmatphys-031214-014710. S2CID 15672131.
- ↑ Zöttl, Andreas; Stark, Holger (11 May 2016). "Emergent behavior in active colloids". Journal of Physics: Condensed Matter. IOP Publishing. 28 (25): 253001. arXiv:1601.06643. Bibcode:2016JPCM...28y3001Z. doi:10.1088/0953-8984/28/25/253001. ISSN 0953-8984. S2CID 3948148.
- ↑ Sutton, Richard (2018). Reinforcement learning : an introduction. Cambridge, Massachusetts: The MIT Press. ISBN 978-0-262-35270-3.
- ↑ Cichos, Frank; Gustavsson, Kristian; Mehlig, Bernhard; Volpe, Giovanni (2020). "Machine learning for active matter". Nature Machine Intelligence. 2 (2): 94–103. doi:10.1038/s42256-020-0146-9. S2CID 214355969.
- ↑ Garnier, Paul; Viquerat, Jonathan; Rabault, Jean; Larcher, Aurélien; Kuhnle, Alexander; Hachem, Elie (2021). "A review on deep reinforcement learning for fluid mechanics". Computers & Fluids. 225: 104973. arXiv:1908.04127. doi:10.1016/j.compfluid.2021.104973. S2CID 199543817.
- ↑ Colabrese, Simona; Gustavsson, Kristian; Celani, Antonio; Biferale, Luca (2017). "Flow Navigation by Smart Microswimmers via Reinforcement Learning". Physical Review Letters. 118 (15): 158004. arXiv:1701.08848. Bibcode:2017PhRvL.118o8004C. doi:10.1103/PhysRevLett.118.158004. PMID 28452499. S2CID 13695532.
- ↑ Yang, Yuguang; Bevan, Michael A.; Li, Bo (2020). "Micro/Nano Motor Navigation and Localization via Deep Reinforcement Learning". Advanced Theory and Simulations. 3 (6). arXiv:2002.06775. doi:10.1002/adts.202000034. S2CID 211133324.
- ↑ Feynman, R. (2018). "There’s plenty of room at the bottom". In: Hey, Anthony (2018). Feynman and computation : exploring the limits of computers. Boca Raton: CRC Press. pp. 63–76. ISBN 978-0-429-50045-9.
- ↑ Kuzajewska, Danuta; Wszołek, Agata; Żwierełło, Wojciech; Kirczuk, Lucyna; Maruszewska, Agnieszka (19 May 2020). "Magnetotactic Bacteria and Magnetosomes as Smart Drug Delivery Systems: A New Weapon on the Battlefield with Cancer?". Biology. MDPI AG. 9 (5): 102. doi:10.3390/biology9050102. ISSN 2079-7737. PMC 7284773. PMID 32438567.
- ↑ Sitti, Metin (2009). "Voyage of the microrobots". Nature. 458 (7242): 1121–1122. doi:10.1038/4581121a. PMID 19407789. S2CID 205044764.
- ↑ Harari, Yuval (2016). Homo deus : a brief history of tomorrow. London: Harvill Secker. ISBN 978-1-4735-4537-3.
- ↑ Qiu, Famin; Fujita, Satoshi; Mhanna, Rami; Zhang, Li; Simona, Benjamin R.; Nelson, Bradley J. (2015). "Magnetic Helical Microswimmers Functionalized with Lipoplexes for Targeted Gene Delivery". Advanced Functional Materials. 25 (11): 1666–1671. doi:10.1002/adfm.201403891. S2CID 95812709.
- ↑ Park, Byung-Wook; Zhuang, Jiang; Yasa, Oncay; Sitti, Metin (2017). "Multifunctional Bacteria-Driven Microswimmers for Targeted Active Drug Delivery". ACS Nano. 11 (9): 8910–8923. doi:10.1021/acsnano.7b03207. PMID 28873304.
- ↑ Wang, Joseph; Gao, Wei (2012). "Nano/Microscale Motors: Biomedical Opportunities and Challenges". ACS Nano. 6 (7): 5745–5751. doi:10.1021/nn3028997. PMID 22770233.
- ↑ Ma, Xing; Hahn, Kersten; Sanchez, Samuel (2015). "Catalytic Mesoporous Janus Nanomotors for Active Cargo Delivery". Journal of the American Chemical Society. 137 (15): 4976–4979. doi:10.1021/jacs.5b02700. PMC 4440854. PMID 25844893.
- ↑ Demirörs, Ahmet F.; Akan, Mehmet Tolga; Poloni, Erik; Studart, André R. (2018). "Active cargo transport with Janus colloidal shuttles using electric and magnetic fields". Soft Matter. 14 (23): 4741–4749. Bibcode:2018SMat...14.4741D. doi:10.1039/C8SM00513C. PMID 29799053.
- ↑ Nelson, Bradley J.; Kaliakatsos, Ioannis K.; Abbott, Jake J. (2010). "Microrobots for Minimally Invasive Medicine". Annual Review of Biomedical Engineering. 12: 55–85. doi:10.1146/annurev-bioeng-010510-103409. PMID 20415589.
- ↑ Soto, Fernando; Wang, Jie; Ahmed, Rajib; Demirci, Utkan (2020). "Medical Micro/Nanorobots in Precision Medicine". Advanced Science. 7 (21). doi:10.1002/advs.202002203. PMC 7610261. PMID 33173743.
- 1 2 Ceylan, Hakan; Yasa, Immihan C; Kilic, Ugur; Hu, Wenqi; Sitti, Metin (16 July 2019). "Translational prospects of untethered medical microrobots". Progress in Biomedical Engineering. IOP Publishing. 1 (1): 012002. doi:10.1088/2516-1091/ab22d5. ISSN 2516-1091. S2CID 199341199.
 Material was copied from this source, which is available under a Creative Commons Attribution 3.0 International License.
Material was copied from this source, which is available under a Creative Commons Attribution 3.0 International License.