RNA-binding protein 4 is a protein that in humans is encoded by the RBM4 gene.[5][6][7]
RBM4 is a protein categorized as an RNA recognition motif (RRM). RRM proteins represent a substantial and functionally varied category of RNA-binding proteins, participating in various functions such as RNA processing and transport, the control of RNA stability, and translational regulation.[8] The RBM4 protein also can be recognized as Lark. Over the past five years, data has emerged from investigations involving mammalian cells, providing a clearer understanding of RBM4's functions. It is now evident that RBM4 serves as an RNA-binding protein, participating in a wide array of cellular processes, which encompass the alternative splicing of pre-mRNA, translation control, and RNA silencing. Structurally, RBM4 shares similarities with other RNA-binding proteins, featuring two RNA recognition motifs and a CCHC-type zinc finger.[9] Notably, RBM4 demonstrates a high degree of conservation throughout the process of evolution. In the context of the human RBM4 gene, it shares an impressive 95% similarity with its murine counterpart.
Structure
Within the realm of mammals, the two distinct isoforms of RBM4 are RBM4a and RBM4b. These two isoforms exhibit remarkably similar sequences, and their existence is believed to have arisen through gene duplication.[10] In humans, both are located on chromosome 11q13, and in mouse are on chromosome 19A.[9] The deletion of both RNA recognition motifs results in anomalous subnuclear localization, indicating that having at least one RRM is essential for directing RBM4 to specific nuclear subcompartments. Also, it appears that RBM4A is not vital for the targeting of RBM4 to the nucleolus. Furthermore, a mutation in RBM4B led to irregular nucleolar expression, extending to its periphery. These observations align with findings from studies on Drosophila Lark, where it was shown that RRM1 is dispensable for embryonic development. In contrast, a double mutation affecting both RNA binding domains is lethal, and a mutation in RRM2 leads to female infertility or developmental arrest in offspring.[11] These two RNA recognition motifs are located at the N-terminus, followed by the CCHC-type zinc finger.
However, the C-terminal region of RBM4 proteins displays less conservation among species. In mammalian RBM4 proteins, there are several stretches rich in alanine, while Drosophila Lark contains sequences that are rich in proline and numerous arginine/serine dipeptides. Previous research has demonstrated the pivotal role of the C-terminal domain of human RBM4 in facilitating its nuclear entry and localization within nuclear speckles enriched with splicing factors. Additionally, this C-terminal region contributes significantly to RBM4's involvement in controlling alternative splicing.
Function
It was initially discovered for its involvement in regulating the circadian rhythm in Drosophila. RBM4 abundance undergoes circadian fluctuations in extracts derived from pharate adults. Additionally, RBM4 is identifiable within neurons that harbor the modulatory neuropeptide crustacean cardioactive peptide (CCAP), playing a crucial role in the regulation of ecdysis in Drosophila and other insects. In CCAP cells, RBM4 exhibits a distinct cytoplasmic localization (as opposed to its nuclear localization in other neurons), and there are striking circadian variations in RBM4 immunoreactivity within this specific neuronal population. Also, RBM4 exhibits widespread expression throughout embryogenesis, with detectable mRNA present in both the developing nervous system and nonneural tissues. Normal embryogenesis necessitates the expression of both maternal and zygotic RBM4. Zygotic RBM4 deficiency halts development around the time of germ band retraction, while embryos lacking maternally inherited RBM4 mRNA experience developmental arrest in early embryogenesis. The retroviral-type zinc finger is crucial for the maternal developmental function, as evidenced by the fact that females with a mutation in this domain can survive and mate, but their zygotes undergo developmental arrest in the early stages of embryogenesis.[12]
RBM4 exhibits dynamic movement between the nucleus and cytoplasm, and this nucleocytoplasmic transport and subcellular positioning are likely under the regulation of cellular signaling pathways. RBM4 undergoes phosphorylation in response to cellular stress conditions and during the initiation of muscle cell differentiation. Phosphorylated RBM4 tends to accumulate in the cytoplasm and can be found in cytoplasmic stress granules as well as granules containing microRNPs (miRNPs). The MKK/p38 kinase pathway is responsible for the stress-induced phosphorylation of RBM4 and its subsequent accumulation in the cytoplasm.[13] Under normal circumstances, RBM4 inhibits Cap-dependent translation. However, when exposed to a stress stimulus, it activates internal ribosome entry site (IRES)-mediated translation, potentially by enhancing the stability of eIF4A-containing initiation complexes. This IRES-mediated translation process facilitates the expression of stress-response genes, suggesting that RBM4 may serve as a translational regulator for stress-associated mRNAs. The role of RBM4 in translation was substantiated by demonstrating its direct binding to the 3'-UTR of Period1 (Per1), a significant circadian clock gene. Murine RBM4 exerted control over the expression of mPER1 in a Cap/poly(A)-dependent manner.[14]
RBM4 can influence alternative 5′-splice site and exon selection in in vivo and in vitro splicing models. By either including or skipping alternative exons, RBM4 demonstrates the dual capacity to function as an activator and a repressor.[15] Notably, RBM4 binds to intronic CU-rich elements in a skeletal muscle-specific isoform of α-tropomyosin, underscoring its interaction with pre-mRNA. Furthermore, RBM4 regulates this isoform in a manner that opposes the actions of polypyrimidine tract binding (PTB) protein, another RNA-binding protein involved in various RNA processing aspects.[16] The competition for binding to the same cis-element is likely between RBM4 and PTB. The binding of RBM4 to intronic elements was corroborated by its interaction with a pyrimidine-rich sequence downstream of the 5′-splice site of exon 10 of tau.
Clinical relevance
Pancreas cell signaling circuit
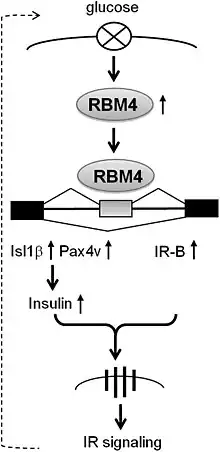
RBM4 plays a role in modulating the inclusion of exon 11 in IR, promoting the expression of the IR-B isoform across various cell lines and tissues. RBM4 functions as a positive regulator in glucose signaling pathways, with its over expression mimicking the effects of glucose induction on insulin and IR-B isoform expression. The prevalence of the IR-B isoform in differentiated pancreatic β cells, exhibiting heightened sensitivity to insulin, is noteworthy. The activation of p38 MAPK signaling by IR-B is crucial for pancreatic development and insulin production in β cells. Additionally, insulin activates the phosphatidylinositol-3-kinase–Akt pathway, supporting β-islet survival and function. Consequently, elevated levels of insulin and IR-B may amplify the signaling pathways of p38 MAPK and Akt. Intriguingly, our findings indicate that RBM4 is responsible for signal-activated p38 MAPK and Akt signaling when cells are exposed to glucose. It is plausible that the increased expression of RBM4 induced by glucose facilitates a more effective transmission of insulin and/or IR signaling, leading to enhanced glucose uptake.[17] Figure 1 displays the framework for a gene expression and signaling circuit in the pancreas regulated by RBM4. RBM4 influences alternative splicing of various pancreatic factors, such as Isl1, Pax4, and IR. The isoforms of Isl1 and Pax4 induced by RBM4 have the potential to enhance insulin gene transcription. Additionally, the IR-B isoform induced by RBM4 likely demonstrates increased signaling activity upon insulin binding, leading to elevated glucose uptake and signaling (represented by the dashed line).[17]
Disease relevance
RBM4 engages in interactions with cyclin A1 and undergoes phosphorylation by cyclin A1–CDK2. The levels of RBM4 and cyclin A1 exhibit a correlation in normal testis, testes with diminished fertility, and in testicular tumors. Cyclin A1 displays tissue-specific expression, particularly high in the testis, where it plays a crucial role in spermatogenesis. Additionally, its presence is noted in specific myeloid leukemias. WT1, known for binding to RBM4, is essential for proper gonadal development and spermatogenesis, and is frequently over expressed in acute myeloid leukemia. Furthermore, reports indicate an up-regulation of RBM4 in apoptotic K562 cells, a human leukemia cell line. In has been see that a decreased amount of RBM4 in the fetal brain results in Down's Syndrome. Considering the expression pattern of RBM4 in the brain and its capability to impact tau splicing, it is plausible that RBM4 may play a role in tauopathies, which encompass a cluster of neurodegenerative disorders, including dementia.[9]
References
- 1 2 3 GRCh38: Ensembl release 89: ENSG00000173933 - Ensembl, May 2017
- 1 2 3 GRCm38: Ensembl release 89: ENSMUSG00000094936 - Ensembl, May 2017
- ↑ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ↑ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ↑ Jackson FR, Banfi S, Guffanti A, Rossi E (May 1997). "A novel zinc finger-containing RNA-binding protein conserved from fruitflies to humans". Genomics. 41 (3): 444–452. doi:10.1006/geno.1997.4704. PMID 9169144.
- ↑ Lin JC, Tarn WY (November 2005). "Exon selection in alpha-tropomyosin mRNA is regulated by the antagonistic action of RBM4 and PTB". Molecular and Cellular Biology. 25 (22): 10111–10121. doi:10.1128/MCB.25.22.10111-10121.2005. PMC 1280272. PMID 16260624.
- ↑ "Entrez Gene: RBM4 RNA binding motif protein 4".
- ↑ Daubner GM, Cléry A, Allain FH (February 2013). "RRM-RNA recognition: NMR or crystallography…and new findings". Current Opinion in Structural Biology. 23 (1): 100–108. doi:10.1016/j.sbi.2012.11.006. PMID 23253355.
- 1 2 3 Markus MA, Morris BJ (April 2009). "RBM4: a multifunctional RNA-binding protein". The International Journal of Biochemistry & Cell Biology. 41 (4): 740–743. doi:10.1016/j.biocel.2008.05.027. PMID 18723113.
- ↑ Zhang P, Wu W, Ma C, Du C, Huang Y, Xu H, et al. (July 2022). "RNA-Binding Proteins in the Regulation of Adipogenesis and Adipose Function". Cells. 11 (15): 2357. doi:10.3390/cells11152357. PMC 9367552. PMID 35954201.
- ↑ Markus MA, Morris BJ (August 2006). "Lark is the splicing factor RBM4 and exhibits unique subnuclear localization properties". DNA and Cell Biology. 25 (8): 457–464. doi:10.1089/dna.2006.25.457. PMID 16907643.
- ↑ McNeil GP, Schroeder AJ, Roberts MA, Jackson FR (September 2001). "Genetic analysis of functional domains within the Drosophila LARK RNA-binding protein". Genetics. 159 (1): 229–240. doi:10.1093/genetics/159.1.229. PMC 1461808. PMID 11560900.
- ↑ Lin JC, Tarn WY (January 2012). "Multiple roles of RBM4 in muscle cell differentiation". Frontiers in Bioscience. 4 (1): 181–189. doi:10.2741/260. PMID 22202052.
- ↑ Kojima S, Matsumoto K, Hirose M, Shimada M, Nagano M, Shigeyoshi Y, et al. (February 2007). "LARK activates posttranscriptional expression of an essential mammalian clock protein, PERIOD1". Proceedings of the National Academy of Sciences of the United States of America. 104 (6): 1859–1864. Bibcode:2007PNAS..104.1859K. doi:10.1073/pnas.0607567104. PMC 1794262. PMID 17264215.
- ↑ Lai MC, Kuo HW, Chang WC, Tarn WY (March 2003). "A novel splicing regulator shares a nuclear import pathway with SR proteins". The EMBO Journal. 22 (6): 1359–1369. doi:10.1093/emboj/cdg126. PMC 151058. PMID 12628928.
- ↑ Lin JC, Tarn WY (November 2005). "Exon selection in alpha-tropomyosin mRNA is regulated by the antagonistic action of RBM4 and PTB". Molecular and Cellular Biology. 25 (22): 10111–10121. doi:10.1128/mcb.25.22.10111-10121.2005. PMC 1280272. PMID 16260624.
- 1 2 3 Lin JC, Yan YT, Hsieh WK, Peng PJ, Su CH, Tarn WY (January 2013). "RBM4 promotes pancreas cell differentiation and insulin expression". Molecular and Cellular Biology. 33 (2): 319–327. doi:10.1128/mcb.01266-12. PMC 3554116. PMID 23129807.
Further reading
- Maruyama K, Sugano S (January 1994). "Oligo-capping: a simple method to replace the cap structure of eukaryotic mRNAs with oligoribonucleotides". Gene. 138 (1–2): 171–174. doi:10.1016/0378-1119(94)90802-8. PMID 8125298.
- Suzuki Y, Yoshitomo-Nakagawa K, Maruyama K, Suyama A, Sugano S (October 1997). "Construction and characterization of a full length-enriched and a 5'-end-enriched cDNA library". Gene. 200 (1–2): 149–156. doi:10.1016/S0378-1119(97)00411-3. PMID 9373149.
- Andersen JS, Lyon CE, Fox AH, Leung AK, Lam YW, Steen H, et al. (January 2002). "Directed proteomic analysis of the human nucleolus". Current Biology. 12 (1): 1–11. doi:10.1016/S0960-9822(01)00650-9. PMID 11790298. S2CID 14132033.
- Bernert G, Fountoulakis M, Lubec G (December 2002). "Manifold decreased protein levels of matrin 3, reduced motor protein HMP and hlark in fetal Down's syndrome brain". Proteomics. 2 (12): 1752–1757. doi:10.1002/1615-9861(200212)2:12<1752::AID-PROT1752>3.0.CO;2-Y. PMID 12469345. S2CID 32160196.
- Petersen HH, Hilpert J, Militz D, Zandler V, Jacobsen C, Roebroek AJ, Willnow TE (February 2003). "Functional interaction of megalin with the megalinbinding protein (MegBP), a novel tetratrico peptide repeat-containing adaptor molecule". Journal of Cell Science. 116 (Pt 3): 453–461. doi:10.1242/jcs.00243. PMID 12508107.
- Lai MC, Kuo HW, Chang WC, Tarn WY (March 2003). "A novel splicing regulator shares a nuclear import pathway with SR proteins". The EMBO Journal. 22 (6): 1359–1369. doi:10.1093/emboj/cdg126. PMC 151058. PMID 12628928.
- Diederichs S, Bäumer N, Ji P, Metzelder SK, Idos GE, Cauvet T, et al. (August 2004). "Identification of interaction partners and substrates of the cyclin A1-CDK2 complex". The Journal of Biological Chemistry. 279 (32): 33727–33741. doi:10.1074/jbc.M401708200. PMID 15159402.
- Rush J, Moritz A, Lee KA, Guo A, Goss VL, Spek EJ, et al. (January 2005). "Immunoaffinity profiling of tyrosine phosphorylation in cancer cells". Nature Biotechnology. 23 (1): 94–101. doi:10.1038/nbt1046. PMID 15592455. S2CID 7200157.
- Rual JF, Venkatesan K, Hao T, Hirozane-Kishikawa T, Dricot A, Li N, et al. (October 2005). "Towards a proteome-scale map of the human protein-protein interaction network". Nature. 437 (7062): 1173–1178. Bibcode:2005Natur.437.1173R. doi:10.1038/nature04209. PMID 16189514. S2CID 4427026.
- Kimura K, Wakamatsu A, Suzuki Y, Ota T, Nishikawa T, Yamashita R, et al. (January 2006). "Diversification of transcriptional modulation: large-scale identification and characterization of putative alternative promoters of human genes". Genome Research. 16 (1): 55–65. doi:10.1101/gr.4039406. PMC 1356129. PMID 16344560.
- Kar A, Havlioglu N, Tarn WY, Wu JY (August 2006). "RBM4 interacts with an intronic element and stimulates tau exon 10 inclusion". The Journal of Biological Chemistry. 281 (34): 24479–24488. doi:10.1074/jbc.M603971200. PMC 2072872. PMID 16777844.
- Markus MA, Heinrich B, Raitskin O, Adams DJ, Mangs H, Goy C, et al. (October 2006). "WT1 interacts with the splicing protein RBM4 and regulates its ability to modulate alternative splicing in vivo". Experimental Cell Research. 312 (17): 3379–3388. doi:10.1016/j.yexcr.2006.07.008. PMID 16934801.
- Lin JC, Hsu M, Tarn WY (February 2007). "Cell stress modulates the function of splicing regulatory protein RBM4 in translation control". Proceedings of the National Academy of Sciences of the United States of America. 104 (7): 2235–2240. Bibcode:2007PNAS..104.2235L. doi:10.1073/pnas.0611015104. PMC 1893002. PMID 17284590.