| STIM2 | |||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| Identifiers | |||||||||||||||||||||||||||||||||||||||||||||||||||
| Aliases | STIM2, stromal interaction molecule 2 | ||||||||||||||||||||||||||||||||||||||||||||||||||
| External IDs | OMIM: 610841 MGI: 2151156 HomoloGene: 32490 GeneCards: STIM2 | ||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| Wikidata | |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
Stromal interaction molecule 2 (STIM2) is a protein that in humans is encoded by the STIM2 gene.[5][6]
This gene is a member of the stromal interaction molecule (STIM) family which comprises only two members together with its homologue STIM1, and likely arose from a common ancestral gene. They encode type 1 transmembrane proteins that are located in the sarco/endoplasmic reticulum (SR / ER) into the cell. Alternative translation initiation from an AUG and a non-AUG (UUG) start site results in the production of two different STIM2 isoforms.
Both members of the STIM family were identified in 2005 as free-calcium (Ca2+) sensors which participate in a mechanism of Ca2+ entry into the cell referred to as store-operated Ca2+ entry (SOCE). Many cellular processes and signaling pathways are started by previous release of Ca2+ stored in subcellular organelles, which needs of a continuous refilling. SOCE is considered the mechanism of store refilling and an essential mechanism of Ca2+ signaling in non-electrically excitable cells. While STIM1 triggers SOCE, research on STIM2 function suggests a major role as feedback regulator that stabilizes basal cytosolic and S/ER Ca2+ concentration [Ca2+]. STIM2 detects small decreases in Ca2+ content stored in the S/ER, switches to the activated state and interacts with so called store-operated Ca2+ (SOC) channels located in the plasma membrane, such as Orai or TRPC channels, allowing SOCE. Although the functional role of STIM2 has been elusive for many years, studies performed in 2009-2010 on murine models suggested that STIM2 participates in processes of the development and functioning of many cell types, including smooth muscle myoblasts, cells of the immune system and neurons, and is involved in tumorigenesis, the development of autoimmune diseases and mechanisms of neuronal damage after transient ischemic conditions.
Gene
In 2001, STIM2 was identified as a new human homologue of the STIM1 gene, representing the second member of a two-gene family in vertebrates.[5] The STIM2 gene contains 12 exons and 11 introns located on the human chromosome 4p15.1, and on the large arm of the mouse chromosome 5, close to the centromere. The members of STIM family most probably have evolved from a single gene in lower multicellular eukaryotes into two related genes in vertebrates, since human STIM1 and STIM2 as well as Drosophila melanogaster Stim (D-Stim) have a conserved genomic organization. The D-STIM protein of 570 aas exhibits equal similarity to both STIM1 (33% identical; 50% of amino acid sequence conserved) and STIM2 (31% identical; 46% of amino acid sequence conserved). Unicellular eukaryotes such as Monosiga brevicollis, a unicellular choanoflagellate has been reported to have a STIM-like gene,[7] however no STIM-like genes have been identified in prokaryotes. No additional STIM-like proteins have been identified until now in vertebrates.[5]
Protein structure
STIM2 protein is a type I transmembrane protein located in the S/ER. Human STIM2 consists of 833 amino acid residues (aas) (105-115 kDa) (Fig. 1), 148 additional aas compared to human STIM1. Their N-terminal regions share 66% similarity over 577 aas (85% of the amino acid sequence of STIM1). Only the extreme of the C-terminal region shows a significant sequence divergence. The domain architecture of both isoforms is highly conserved in vertebrates (Fig. 1). Mouse STIM2 shares a 92% identity with human STIM2 in the aminoacid sequence according to the pairwise alignment generated by BLAST. Their domain structure is also highly conserved (Fig. 1). Human STIM2 is post-translationally modified in vivo, such as maturation by cleavage of N-terminal S/ER signaling peptide (14 aas), glycosylation and variable degrees of phosphorylation, but the phosphorylated sites are still unknown (Fig. 1).[5]
Domain architecture
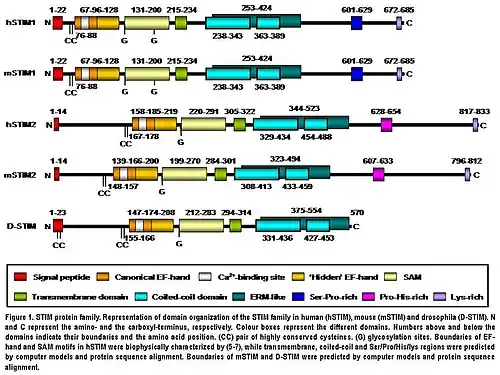
The N-terminal region of STIM2 is located in the S/ER lumen and contains a canonical EF-hand Ca2+-binding motif, a “hidden” EF-hand Ca2+-binding motif discovered recently and a sterile a-motif (SAM) domain, a well-known protein–protein interaction motif (Fig. 1).[8][9][10] The N-terminal portion is separated from the C-terminal region by a single-pass transmembrane motif that is highly conserved in all STIM proteins. The C-terminal region contains a high degree of α-helical structures. A large proportion close to the transmembrane domain comprises a region similar to an ezrin/radixin/moesin (ERM) domain that contains two coiled-coil domains.[11] The coiled-coil domains mediate interactions between STIM proteins, allowing them to bind each other and form homo and heterodimers (Fig. 1).[12][13][14] Finally, further towards the C-terminus, STIM2 contains a proline/histidine-rich motif and a lysine-rich tail of 17 aas (Fig. 1).[5]
EF-hand-SAM region
Since the EF-hand and SAM (EF-SAM) domains are vital to STIM function and SOCE regulation, they are now discussed in detail. The EF-hand domain is a Ca2+ sensor used by STIM protein to detect changes in Ca2+ concentration inside the S/ER. STIM isoforms become activated when Ca2+ bound to the EF-hand motif is released as a result of a decrease in Ca2+ levels inside the S/ER store after IP3 receptor–mediated depletion. It has been reported that STIM EF-hand mutants that are not able to bind Ca2+ are constitutively active and continually activate SOCE independently of S/ER [Ca2+], in vitro[15] and in vivo.[16][17][18] The SAM domain is important for STIM oligomerization, since mutants in this domain lack the ability to form inducible punctae.[19] Ca2+-binding experiments in vitro using human STIM1 EF–SAM (residue 58–201) or STIM2 EF–SAM (residue 149–292) fragments show that both isoforms bind Ca2+ with similar affinity (STIM2 Kd~0.5 mM; STIM1 Kd~0.2–0.6 mM),[20][21] which is within the range of values reported for S/ER [Ca2+].[22][23] However, STIM2 differs from STIM1 in that it is already partially active at basal S/ER [Ca2+] and becomes fully activated earlier during S/ER store depletion. Despite the same Ca2+ affinity shown by STIM EF-SAM fragments, the full STIM2 protein showed a lower [Ca2+] sensitivity than STIM1 in transfected cells in vitro.[24] This discrepancy indicates that other protein regions in addition contribute to the different [Ca2+] sensitivity or activation threshold shown by both isoforms. The “hidden” EF-hand domain does not bind Ca2+, but it is critical for intramolecular association, folding and stability of the EF-hand and SAM domains. Very recently it has been reported that structurally critical mutations in the canonical EF-hand, ‘‘hidden’’ EF-hand, or SAM domain disrupt Ca2+ sensitivity due to the destabilization of the entire EF-SAM region.[25]
C-terminal region
Besides the N-terminus, the C-terminal region is also an essential part of STIM proteins. It shows a significant sequence divergence between both isoforms and in STIM1, the C-terminal region is essential for the interaction with SOC channels.[26] Human STIM2 contains a proline- and histidine-rich motif (PHAPHPSHPRHPHHPQHTPHSLPSPDP) at a similar position to a serine- and proline-rich region (SPSAPPGGSPHLDSSRSHSPSSPDPDTPSP) in STIM1. The significant divergence in these regions could indicate a divergence in function of STIM isoforms. Unlike STIM1, STIM2 has a dilysine ER retention signal (K(X)KXX) at its extreme C-terminus which retains the protein in the ER, whereas STIM1 can travel to cell surface.[27] Finally, similar lysine-rich tails of 14 and 17 residues in STIM1 and STIM2 respectively are located at the very end of the C-terminal region. Linear peptides from C-terminal polybasic region of human STIM1 (residues 667-685) and STIM2 (residues 730-746) bind to calmodulin with high or low affinity in presence or absence of Ca2+, respectively.[28] Most of studies on interactions of the C-terminal region have been performed with the STIM1 isoform. The addition of thapsigargin (the SERCA pump inhibitor that stimulates SOCE by passive depletion of intracellular Ca2+ stores) to human salivary gland cells as well as dispersed mouse submandibular gland cells increase coimmunoprecipitation of TRPC1 and Orai1 with STIM1.[29] By in vitro co-expression of different human STIM1 mutants that lack the different C-terminal regions in HEK293 cells, three recent works reported that the ERM domain in the C-terminus (aas 251-535, Fig. 1), containing the coiled-coil domains, mediates the binding of STIM1 to TRPC(1, 2,4 and 5) and the STIM1 migration to the plasma membrane. Furthermore, the cationic lysine-rich region is essential for gating of TRPC1.[14][26][30] Li et al. further delineated these regions (aas 425-672) as possible STIM1-Orai1 interaction sites.[14] In vitro coimmunoprecipitation experiments after transient coexpression of STIM2 and Orai1 proteins in HEK293 cells revealed that also STIM2 can physically interact with Orai1, probably though the STIM2 C-terminal region.[31]
Expression and tissue distribution
STIM2 mRNA is expressed by most human tissues. The STIM2 protein is expressed by many human cell lines together with STIM1, indicating that STIM isoforms are co-expressed in the same cell, at least in the established cell lines.[5] STIM2 protein is widely expressed in tissues, usually present at lower levels than STIM1 except in brain or liver, were STIM2 seems to be the dominant isoform.[5][32] Stim2 transcription is also dynamically regulated, for instance being upregulated upon differentiation of naive T cells into Th1 or Th2 lymphocytes.[33]
Function
The STIM2 function has been controversial. Initial studies found that siRNA knockdown of STIM1, but not STIM2, strongly reduced SOCE in mammalian cells.[15][24][34][35][36] Liou et al. reported a slight reduction in SOCE also by knockdown of STIM2 in HeLa cells.[15] Soboloff et al. suggested that STIM2 inhibits SOCE when expressed alone,[13] but coexpressed with Orai1 causes substantial constitutive SOCE.[37] In contrast, Brandman et al. suggested that STIM2 could act as a regulator that stabilizes basal cytosolic and ER Ca2+ levels.[24] Parvez et al., using in vitro transient coexpression of human STIM2 and different SOC channels in HEK293 cells, reported that STIM2 mediates SOCE via two store-dependent and store independent modes.[31] Taking together, these results indicate a complex interaction finely regulated by the STIM1: STIM2: Orai cellular ratio and their endogenous levels.
Studies performed in 2009-2010 using human in vitro or murine in vivo models confirmed Brandman et al. results and suggested that STIM2 participates in processes of the development and functioning of many cell types, including smooth muscle myoblasts, cells of the immune system and neurons. Moreover, it is involved in tumorigenesis, the development of autoimmune diseases and mechanisms of neuronal damage after transient ischemic conditions. In resting conditions, cultured HEK293 cells overexpressing or cortical neurons lacking STIM2 have increased or decreased resting intracellular Ca2+ levels respectively,[31][32] supporting the idea that STIM2 is essential for regulation of intracellular basal Ca2+ levels. However, cells are very active in vivo and intracellular Ca2+ levels are continuously fluctuating. The development of new methods to study the in vivo role of STIM2 in intracellular Ca2+ levels would be necessary. In cultured human myoblast, STIM2 participate in cell differentiation into myotubes.[38] In the immune system, STIM2 participates in T cell activation-induced production of interleukin2 (IL-2) and interferon gamma (IFNγ), probably by stabilization of NFAT residence in the nucleus, as well as in differentiation of naive T cells into Th17 lymphocytes, which presumably are important in early phases of autoimmune diseases.[33][39] In fact, STIM2-deficient mice showed mild symptomatology in the early phase of autoimmune diseases.[39] In neuronal tissue, STIM2 plays a crucial role in ischemia-induced neuronal damage, and the absence of STIM2 in knockout mice reduced the neuronal damage produced by ischemia after transient interruption of blood flow in brain.[32] This neuroprotective effect of STIM2-deficiency after an ischemic episode indicates that inhibitors of STIM2 function may thus have a potential therapeutic value as neuroprotective agents to treat ischemic injury and other neurodegenerative disorders involving altered Ca2+ homeostasis. Moreover, the same scientific study suggested an important role of STIM2 in hippocampus-dependent spatial memory, synaptic transmission and plasticity.[32]
Finally, an oncogenic function has been demonstrated for STIM2, together with STIM1, in glioblastoma multiforme, where both proteins have increased expression and/or increased copy number.[40][41] Additionally, STIM2 is located in chromosome 4p15.1, a region implicated in invasive carcinomas of the lung, breast, neck and head.[42][43][44]
Interactions
As mentioned before, STIM2 has been shown to interact with STIM1,[12][13][14] SOC channels such as Orai (ICRACM) or TRPC,[31] calmodulin (CaM)[28][31] and also plasma membrane phosphoinositides.[45] The expression of STIM2 has been shown to be influenced or regulated by presenilins in mouse embryonic fibroblasts and human B lymphocytes.[46]
References
- 1 2 3 GRCh38: Ensembl release 89: ENSG00000109689 - Ensembl, May 2017
- 1 2 3 GRCm38: Ensembl release 89: ENSMUSG00000039156 - Ensembl, May 2017
- ↑ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ↑ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- 1 2 3 4 5 6 7 Williams RT, Manji SS, Parker NJ, Hancock MS, Van Stekelenburg L, Eid JP, Senior PV, Kazenwadel JS, Shandala T, Saint R, Smith PJ, Dziadek MA (August 2001). "Identification and characterization of the STIM (stromal interaction molecule) gene family: coding for a novel class of transmembrane proteins". The Biochemical Journal. 357 (Pt 3): 673–85. doi:10.1042/0264-6021:3570673. PMC 1221997. PMID 11463338.
- ↑ "Entrez Gene: STIM2 stromal interaction molecule 2".
- ↑ Cai X (July 2008). "Unicellular Ca2+ signaling 'toolkit' at the origin of metazoa". Molecular Biology and Evolution. 25 (7): 1357–61. doi:10.1093/molbev/msn077. PMID 18385221.
- ↑ Kim CA, Bowie JU (December 2003). "SAM domains: uniform structure, diversity of function". Trends in Biochemical Sciences. 28 (12): 625–8. doi:10.1016/j.tibs.2003.11.001. PMID 14659692.
- ↑ Qiao F, Bowie JU (May 2005). "The many faces of SAM". Science's STKE. 2005 (286): re7. doi:10.1126/stke.2862005re7. PMID 15928333. S2CID 31699798.
- ↑ Schultz J, Ponting CP, Hofmann K, Bork P (January 1997). "SAM as a protein interaction domain involved in developmental regulation". Protein Science. 6 (1): 249–53. doi:10.1002/pro.5560060128. PMC 2143507. PMID 9007998.
- ↑ Parry DA, Fraser RD, Squire JM (September 2008). "Fifty years of coiled-coils and alpha-helical bundles: a close relationship between sequence and structure". Journal of Structural Biology. 163 (3): 258–69. doi:10.1016/j.jsb.2008.01.016. PMID 18342539.
- 1 2 Williams RT, Senior PV, Van Stekelenburg L, Layton JE, Smith PJ, Dziadek MA (April 2002). "Stromal interaction molecule 1 (STIM1), a transmembrane protein with growth suppressor activity, contains an extracellular SAM domain modified by N-linked glycosylation". Biochimica et Biophysica Acta (BBA) - Protein Structure and Molecular Enzymology. 1596 (1): 131–7. doi:10.1016/S0167-4838(02)00211-X. PMID 11983428.
- 1 2 3 Soboloff J, Spassova MA, Hewavitharana T, He LP, Xu W, Johnstone LS, Dziadek MA, Gill DL (July 2006). "STIM2 is an inhibitor of STIM1-mediated store-operated Ca2+ Entry". Current Biology. 16 (14): 1465–70. doi:10.1016/j.cub.2006.05.051. PMID 16860747. S2CID 16570108.
- 1 2 3 4 Li Z, Lu J, Xu P, Xie X, Chen L, Xu T (October 2007). "Mapping the interacting domains of STIM1 and Orai1 in Ca2+ release-activated Ca2+ channel activation". The Journal of Biological Chemistry. 282 (40): 29448–56. doi:10.1074/jbc.M703573200. PMID 17702753.
- 1 2 3 Liou J, Kim ML, Heo WD, Jones JT, Myers JW, Ferrell JE, Meyer T (July 2005). "STIM is a Ca2+ sensor essential for Ca2+-store-depletion-triggered Ca2+ influx". Current Biology. 15 (13): 1235–41. doi:10.1016/j.cub.2005.05.055. PMC 3186072. PMID 16005298.
- ↑ Grosse J, Braun A, Varga-Szabo D, Beyersdorf N, Schneider B, Zeitlmann L, Hanke P, Schropp P, Mühlstedt S, Zorn C, Huber M, Schmittwolf C, Jagla W, Yu P, Kerkau T, Schulze H, Nehls M, Nieswandt B (November 2007). "An EF hand mutation in Stim1 causes premature platelet activation and bleeding in mice". The Journal of Clinical Investigation. 117 (11): 3540–50. doi:10.1172/JCI32312. PMC 2040319. PMID 17965774.
- ↑ Mercer JC, Dehaven WI, Smyth JT, Wedel B, Boyles RR, Bird GS, Putney JW (August 2006). "Large store-operated calcium selective currents due to co-expression of Orai1 or Orai2 with the intracellular calcium sensor, Stim1". The Journal of Biological Chemistry. 281 (34): 24979–90. doi:10.1074/jbc.M604589200. PMC 1633822. PMID 16807233.
- ↑ Zhang SL, Yeromin AV, Zhang XH, Yu Y, Safrina O, Penna A, Roos J, Stauderman KA, Cahalan MD (June 2006). "Genome-wide RNAi screen of Ca(2+) influx identifies genes that regulate Ca(2+) release-activated Ca(2+) channel activity". Proceedings of the National Academy of Sciences of the United States of America. 103 (24): 9357–62. doi:10.1073/pnas.0603161103. PMC 1482614. PMID 16751269.
- ↑ Baba Y, Hayashi K, Fujii Y, Mizushima A, Watarai H, Wakamori M, Numaga T, Mori Y, Iino M, Hikida M, Kurosaki T (November 2006). "Coupling of STIM1 to store-operated Ca2+ entry through its constitutive and inducible movement in the endoplasmic reticulum". Proceedings of the National Academy of Sciences of the United States of America. 103 (45): 16704–9. Bibcode:2006PNAS..10316704B. doi:10.1073/pnas.0608358103. PMC 1636519. PMID 17075073.
- ↑ Zheng L, Stathopulos PB, Li GY, Ikura M (April 2008). "Biophysical characterization of the EF-hand and SAM domain containing Ca2+ sensory region of STIM1 and STIM2". Biochemical and Biophysical Research Communications. 369 (1): 240–6. doi:10.1016/j.bbrc.2007.12.129. PMID 18166150.
- ↑ Stathopulos PB, Li GY, Plevin MJ, Ames JB, Ikura M (November 2006). "Stored Ca2+ depletion-induced oligomerization of stromal interaction molecule 1 (STIM1) via the EF-SAM region: An initiation mechanism for capacitive Ca2+ entry". The Journal of Biological Chemistry. 281 (47): 35855–62. doi:10.1074/jbc.M608247200. PMID 17020874.
- ↑ Demaurex N, Frieden M (August 2003). "Measurements of the free luminal ER Ca(2+) concentration with targeted "cameleon" fluorescent proteins". Cell Calcium. 34 (2): 109–19. doi:10.1016/S0143-4160(03)00081-2. PMID 12810053.
- ↑ Barrero MJ, Montero M, Alvarez J (October 1997). "Dynamics of [Ca2+] in the endoplasmic reticulum and cytoplasm of intact HeLa cells. A comparative study". The Journal of Biological Chemistry. 272 (44): 27694–9. doi:10.1074/jbc.272.44.27694. PMID 9346910.
- 1 2 3 Brandman O, Liou J, Park WS, Meyer T (December 2007). "STIM2 is a feedback regulator that stabilizes basal cytosolic and endoplasmic reticulum Ca2+ levels". Cell. 131 (7): 1327–39. doi:10.1016/j.cell.2007.11.039. PMC 2680164. PMID 18160041.
- ↑ Stathopulos PB, Zheng L, Li GY, Plevin MJ, Ikura M (October 2008). "Structural and mechanistic insights into STIM1-mediated initiation of store-operated calcium entry". Cell. 135 (1): 110–22. doi:10.1016/j.cell.2008.08.006. PMID 18854159. S2CID 15447873.
- 1 2 Huang GN, Zeng W, Kim JY, Yuan JP, Han L, Muallem S, Worley PF (September 2006). "STIM1 carboxyl-terminus activates native SOC, I(crac) and TRPC1 channels". Nature Cell Biology. 8 (9): 1003–10. doi:10.1038/ncb1454. PMID 16906149. S2CID 22135544.
- ↑ Ercan E, Chung SH, Bhardwaj R, Seedorf M (July 2012). "Di-arginine signals and the K-rich domain retain the Ca²⁺ sensor STIM1 in the endoplasmic reticulum". Traffic. 13 (7): 992–1003. doi:10.1111/j.1600-0854.2012.01359.x. PMID 22498042. S2CID 33126543.
- 1 2 Bauer MC, O'Connell D, Cahill DJ, Linse S (June 2008). "Calmodulin binding to the polybasic C-termini of STIM proteins involved in store-operated calcium entry". Biochemistry. 47 (23): 6089–91. doi:10.1021/bi800496a. PMID 18484746.
- ↑ Ong HL, Cheng KT, Liu X, Bandyopadhyay BC, Paria BC, Soboloff J, Pani B, Gwack Y, Srikanth S, Singh BB, Gill DL, Gill D, Ambudkar IS (March 2007). "Dynamic assembly of TRPC1-STIM1-Orai1 ternary complex is involved in store-operated calcium influx. Evidence for similarities in store-operated and calcium release-activated calcium channel components". The Journal of Biological Chemistry. 282 (12): 9105–16. doi:10.1074/jbc.M608942200. PMC 3309402. PMID 17224452.
- ↑ Yuan JP, Zeng W, Huang GN, Worley PF, Muallem S (June 2007). "STIM1 heteromultimerizes TRPC channels to determine their function as store-operated channels". Nature Cell Biology. 9 (6): 636–45. doi:10.1038/ncb1590. PMC 2699187. PMID 17486119.
- 1 2 3 4 5 Parvez S, Beck A, Peinelt C, Soboloff J, Lis A, Monteilh-Zoller M, Gill DL, Fleig A, Penner R (March 2008). "STIM2 protein mediates distinct store-dependent and store-independent modes of CRAC channel activation". FASEB Journal. 22 (3): 752–61. doi:10.1096/fj.07-9449com. PMC 3601890. PMID 17905723.
- 1 2 3 4 Berna-Erro A, Braun A, Kraft R, Kleinschnitz C, Schuhmann MK, Stegner D, Wultsch T, Eilers J, Meuth SG, Stoll G, Nieswandt B (2009). "STIM2 regulates capacitive Ca2+ entry in neurons and plays a key role in hypoxic neuronal cell death". Science Signaling. 2 (93): ra67. doi:10.1126/scisignal.2000522. PMID 19843959.
- 1 2 Oh-Hora M, Yamashita M, Hogan PG, Sharma S, Lamperti E, Chung W, Prakriya M, Feske S, Rao A (April 2008). "Dual functions for the endoplasmic reticulum calcium sensors STIM1 and STIM2 in T cell activation and tolerance". Nature Immunology. 9 (4): 432–43. doi:10.1038/ni1574. PMC 2737533. PMID 18327260.
- ↑ Roos J, DiGregorio PJ, Yeromin AV, Ohlsen K, Lioudyno M, Zhang S, Safrina O, Kozak JA, Wagner SL, Cahalan MD, Veliçelebi G, Stauderman KA (May 2005). "STIM1, an essential and conserved component of store-operated Ca2+ channel function". The Journal of Cell Biology. 169 (3): 435–45. doi:10.1083/jcb.200502019. PMC 2171946. PMID 15866891.
- ↑ Zhang SL, Yu Y, Roos J, Kozak JA, Deerinck TJ, Ellisman MH, Stauderman KA, Cahalan MD (October 2005). "STIM1 is a Ca2+ sensor that activates CRAC channels and migrates from the Ca2+ store to the plasma membrane". Nature. 437 (7060): 902–5. Bibcode:2005Natur.437..902Z. doi:10.1038/nature04147. PMC 1618826. PMID 16208375.
- ↑ Spassova MA, Soboloff J, He LP, Xu W, Dziadek MA, Gill DL (March 2006). "STIM1 has a plasma membrane role in the activation of store-operated Ca(2+) channels". Proceedings of the National Academy of Sciences of the United States of America. 103 (11): 4040–5. Bibcode:2006PNAS..103.4040S. doi:10.1073/pnas.0510050103. PMC 1449642. PMID 16537481.
- ↑ Soboloff J, Spassova MA, Tang XD, Hewavitharana T, Xu W, Gill DL (July 2006). "Orai1 and STIM reconstitute store-operated calcium channel function". The Journal of Biological Chemistry. 281 (30): 20661–5. doi:10.1074/jbc.C600126200. PMID 16766533.
- ↑ Darbellay B, Arnaudeau S, Ceroni D, Bader CR, Konig S, Bernheim L (July 2010). "Human muscle economy myoblast differentiation and excitation-contraction coupling use the same molecular partners, STIM1 and STIM2". The Journal of Biological Chemistry. 285 (29): 22437–47. doi:10.1074/jbc.M110.118984. PMC 2903423. PMID 20436167.
- 1 2 Schuhmann MK, Stegner D, Berna-Erro A, Bittner S, Braun A, Kleinschnitz C, Stoll G, Wiendl H, Meuth SG, Nieswandt B (February 2010). "Stromal interaction molecules 1 and 2 are key regulators of autoreactive T cell activation in murine autoimmune central nervous system inflammation". Journal of Immunology. 184 (3): 1536–42. doi:10.4049/jimmunol.0902161. PMID 20028655.
- ↑ Scrideli CA, Carlotti CG, Okamoto OK, Andrade VS, Cortez MA, Motta FJ, Lucio-Eterovic AK, Neder L, Rosemberg S, Oba-Shinjo SM, Marie SK, Tone LG (July 2008). "Gene expression profile analysis of primary glioblastomas and non-neoplastic brain tissue: identification of potential target genes by oligonucleotide microarray and real-time quantitative PCR". Journal of Neuro-Oncology. 88 (3): 281–91. doi:10.1007/s11060-008-9579-4. PMID 18398573. S2CID 31862636.
- ↑ Ruano Y, Mollejo M, Ribalta T, Fiaño C, Camacho FI, Gómez E, de Lope AR, Hernández-Moneo JL, Martínez P, Meléndez B (2006). "Identification of novel candidate target genes in amplicons of Glioblastoma multiforme tumors detected by expression and CGH microarray profiling". Molecular Cancer. 5: 39. doi:10.1186/1476-4598-5-39. PMC 1592108. PMID 17002787.
- ↑ Pershouse MA, Ligon AH, Pereira-Smith OM, Killary AM, Yung WK, Steck PA (November 1997). "Suppression of transformed phenotype and tumorigenicity after transfer of chromosome 4 into U251 human glioma cells". Genes, Chromosomes & Cancer. 20 (3): 260–7. doi:10.1002/(SICI)1098-2264(199711)20:3<260::AID-GCC6>3.0.CO;2-0. PMID 9365833. S2CID 13411377.
- ↑ Richard F, Pacyna-Gengelbach M, Schlüns K, Fleige B, Winzer KJ, Szymas J, Dietel M, Petersen I, Schwendel A (May 2000). "Patterns of chromosomal imbalances in invasive breast cancer". International Journal of Cancer. 89 (3): 305–10. doi:10.1002/1097-0215(20000520)89:3<305::AID-IJC15>3.0.CO;2-8. PMID 10861509.
- ↑ Petersen S, Aninat-Meyer M, Schlüns K, Gellert K, Dietel M, Petersen I (January 2000). "Chromosomal alterations in the clonal evolution to the metastatic stage of squamous cell carcinomas of the lung". British Journal of Cancer. 82 (1): 65–73. doi:10.1054/bjoc.1999.0878. PMC 2363206. PMID 10638968.
- ↑ Ercan E, Momburg F, Engel U, Temmerman K, Nickel W, Seedorf M (December 2009). "A conserved, lipid-mediated sorting mechanism of yeast Ist2 and mammalian STIM proteins to the peripheral ER". Traffic. 10 (12): 1802–18. doi:10.1111/j.1600-0854.2009.00995.x. PMID 19845919. S2CID 31387679.
- ↑ Bojarski L, Pomorski P, Szybinska A, Drab M, Skibinska-Kijek A, Gruszczynska-Biegala J, Kuznicki J (June 2009). "Presenilin-dependent expression of STIM proteins and dysregulation of capacitative Ca2+ entry in familial Alzheimer's disease". Biochimica et Biophysica Acta (BBA) - Molecular Cell Research. 1793 (6): 1050–7. doi:10.1016/j.bbamcr.2008.11.008. PMID 19111578.