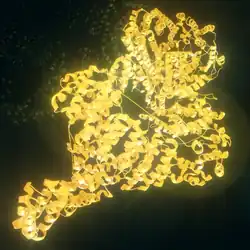
ATM serine/threonine kinase or Ataxia-telangiectasia mutated, symbol ATM, is a serine/threonine protein kinase that is recruited and activated by DNA double-strand breaks (canonical pathway), oxidative stress, topoisomerase cleavage complexes, splicing intermediates, R-loops and in some cases by single-strand DNA breaks.[5] It phosphorylates several key proteins that initiate activation of the DNA damage checkpoint, leading to cell cycle arrest, DNA repair or apoptosis. Several of these targets, including p53, CHK2, BRCA1, NBS1 and H2AX are tumor suppressors.
In 1995, the gene was discovered by Yosef Shiloh[6] who named its product ATM since he found that its mutations are responsible for the disorder ataxia–telangiectasia.[7] In 1998, the Shiloh and Kastan laboratories independently showed that ATM is a protein kinase whose activity is enhanced by DNA damage.[8][9]
Introduction
Throughout the cell cycle DNA is monitored for damage. Damages result from errors during replication, by-products of metabolism, general toxic drugs or ionizing radiation. The cell cycle has different DNA damage checkpoints, which inhibit the next or maintain the current cell cycle step. There are two main checkpoints, the G1/S and the G2/M, during the cell cycle, which preserve correct progression. ATM plays a role in cell cycle delay after DNA damage, especially after double-strand breaks (DSBs).[10] ATM is recruited to sites of double strand breaks by DSB sensor proteins, such as the MRN complex. After being recruited, it phosphorylates NBS1, along other DSB repair proteins. These modified mediator proteins then amplify the DNA damage signal, and transduce the signals to downstream effectors such as CHK2 and p53.
Structure
The ATM gene codes for a 350 kDa protein consisting of 3056 amino acids.[11] ATM belongs to the superfamily of phosphatidylinositol 3-kinase-related kinases (PIKKs). The PIKK superfamily comprises six Ser/Thr-protein kinases that show a sequence similarity to phosphatidylinositol 3-kinases (PI3Ks). This protein kinase family includes ATR (ATM- and RAD3-related), DNA-PKcs (DNA-dependent protein kinase catalytic subunit) and mTOR (mammalian target of rapamycin). Characteristic for ATM are five domains. These are from N-terminus to C-terminus the HEAT repeat domain, the FRAP-ATM-TRRAP (FAT) domain, the kinase domain (KD), the PIKK-regulatory domain (PRD) and the FAT-C-terminal (FATC) domain. The HEAT repeats directly bind to the C-terminus of NBS1. The FAT domain interacts with ATM's kinase domain to stabilize the C-terminus region of ATM itself. The KD domain resumes kinase activity, while the PRD and the FATC domain regulate it. The structure of ATM has been solved in several publications using cryo-EM. In the inactive form, the protein forms a homodimer. In the canonical pathway, ATM is activated by the MRN complex and autophosphorylation, forming active monomers capable of phosphorylating several hundred downstream targets. In the non-canonical pathway, e.g. through simulation by oxidative stress, the dimer can be activated by the formation of disulfide bonds.[12] The entire N-terminal domain together with the FAT domain are adopt an α-helical structure, which was initially predicted by sequence analysis. This α-helical structure forms a tertiary structure, which has a curved, tubular shape present for example in the Huntingtin protein, which also contains HEAT repeats. FATC is the C-terminal domain with a length of about 30 amino acids. It is highly conserved and consists of an α-helix.[13]

Function
A complex of the three proteins MRE11, RAD50 and NBS1 (XRS2 in yeast), called the MRN complex in humans, recruits ATM to double strand breaks (DSBs) and holds the two ends together. ATM directly interacts with the NBS1 subunit and phosphorylates the histone variant H2AX on Ser139.[14] This phosphorylation generates binding sites for adaptor proteins with a BRCT domain. These adaptor proteins then recruit different factors including the effector protein kinase CHK2 and the tumor suppressor p53. The ATM-mediated DNA damage response consists of a rapid and a delayed response. The effector kinase CHK2 is phosphorylated and thereby activated by ATM. Activated CHK2 phosphorylates phosphatase CDC25A, which is degraded thereupon and can no longer dephosphorylate CDK1-cyclin B, resulting in cell-cycle arrest. If the DSB can not be repaired during this rapid response, ATM additionally phosphorylates MDM2 and p53 at Ser15.[9] p53 is also phosphorylated by the effector kinase CHK2. These phosphorylation events lead to stabilization and activation of p53 and subsequent transcription of numerous p53 target genes including CDK inhibitor p21 which lead to long-term cell-cycle arrest or even apoptosis.[15]
.png.webp)
The protein kinase ATM may also be involved in mitochondrial homeostasis, as a regulator of mitochondrial autophagy (mitophagy) whereby old, dysfunctional mitochondria are removed.[16] Increased ATM activity also occurs in viral infection where ATM is activated early during dengue virus infection as part of autophagy induction and ER stress response.[17]
Regulation
A functional MRN complex is required for ATM activation after DSBs. The complex functions upstream of ATM in mammalian cells and induces conformational changes that facilitate an increase in the affinity of ATM towards its substrates, such as CHK2 and p53.[10] Inactive ATM is present in the cells without DSBs as dimers or multimers. Upon DNA damage, ATM autophosphorylates on residue Ser1981. This phosphorylation provokes dissociation of ATM dimers, which is followed by the release of active ATM monomers.[18] Further autophosphorylation (of residues Ser367 and Ser1893) is required for normal activity of the ATM kinase. Activation of ATM by the MRN complex is preceded by at least two steps, i.e. recruitment of ATM to DSB ends by the mediator of DNA damage checkpoint protein 1 (MDC1) which binds to MRE11, and the subsequent stimulation of kinase activity with the NBS1 C-terminus. The three domains FAT, PRD and FATC are all involved in regulating the activity of the KD kinase domain. The FAT domain interacts with ATM's KD domain to stabilize the C-terminus region of ATM itself. The FATC domain is critical for kinase activity and highly sensitive to mutagenesis. It mediates protein-protein interaction for example with the histone acetyltransferase TIP60 (HIV-1 Tat interacting protein 60 kDa), which acetylates ATM on residue Lys3016. The acetylation occurs in the C-terminal half of the PRD domain and is required for ATM kinase activation and for its conversion into monomers. While deletion of the entire PRD domain abolishes the kinase activity of ATM, specific small deletions show no effect.[13]
Germline mutations and cancer risk
People who carry a heterozygous ATM mutation have increased risk of mainly pancreatic cancer, prostate cancer, stomach cancer and invasive ductal carcinoma of the breast.[19] Homozygous ATM mutation confers the disease ataxia–telangiectasia (AT), a rare human disease characterized by cerebellar degeneration, extreme cellular sensitivity to radiation and a predisposition to cancer. All AT patients contain mutations in the ATM gene. Most other AT-like disorders are defective in genes encoding the MRN protein complex. One feature of the ATM protein is its rapid increase in kinase activity immediately following double-strand break formation.[20][8] The phenotypic manifestation of AT is due to the broad range of substrates for the ATM kinase, involving DNA repair, apoptosis, G1/S, intra-S checkpoint and G2/M checkpoints, gene regulation, translation initiation, and telomere maintenance.[21] Therefore, a defect in ATM has severe consequences in repairing certain types of damage to DNA, and cancer may result from improper repair. AT patients have an increased risk for breast cancer that has been ascribed to ATM's interaction and phosphorylation of BRCA1 and its associated proteins following DNA damage.[22]
Somatic ATM mutations in sporadic cancers
Mutations in the ATM gene are found at relatively low frequencies in sporadic cancers. According to COSMIC, the Catalogue Of Somatic Mutations In Cancer, the frequencies with which heterozygous mutations in ATM are found in common cancers include 0.7% in 713 ovarian cancers, 0.9% in central nervous system cancers, 1.9% in 1,120 breast cancers, 2.1% in 847 kidney cancers, 4.6% in colon cancers, 7.2% among 1,040 lung cancers and 11.1% in 1790 hematopoietic and lymphoid tissue cancers.[23] Certain kinds of leukemias and lymphomas, including mantle cell lymphoma, T-ALL, atypical B cell chronic lymphocytic leukemia, and T-PLL are also associated with ATM defects.[24] A comprehensive literature search on ATM deficiency in pancreatic cancer, that captured 5,234 patients, estimated that the total prevalence of germline or somatic ATM mutations in pancreatic cancer was 6.4%.[25] ATM mutations may serve as predictive biomarkers of response for certain therapies, since preclinical studies have found that ATM deficiency can sensitise some cancer types to ATR inhibition.[26][27][28][29]
Frequent epigenetic deficiencies of ATM in cancers
ATM is one of the DNA repair genes frequently hypermethylated in its promoter region in various cancers (see table of such genes in Cancer epigenetics). The promoter methylation of ATM causes reduced protein or mRNA expression of ATM.
More than 73% of brain tumors were found to be methylated in the ATM gene promoter and there was strong inverse correlation between ATM promoter methylation and its protein expression (p < 0.001).[30]
The ATM gene promoter was observed to be hypermethylated in 53% of small (impalpable) breast cancers[31] and was hypermethylated in 78% of stage II or greater breast cancers with a highly significant correlation (P = 0.0006) between reduced ATM mRNA abundance and aberrant methylation of the ATM gene promoter.[32]
In non-small cell lung cancer (NSCLC), the ATM promoter methylation status of paired tumors and surrounding histologically uninvolved lung tissue was found to be 69% and 59%, respectively. However, in more advanced NSCLC the frequency of ATM promoter methylation was lower at 22%.[33] The finding of ATM promoter methylation in surrounding histologically uninvolved lung tissue suggests that ATM deficiency may be present early in a field defect leading to progression to NSCLC.
In squamous cell carcinoma of the head and neck, 42% of tumors displayed ATM promoter methylation.[34]
DNA damage appears to be the primary underlying cause of cancer,[35] and deficiencies in DNA repair likely underlie many forms of cancer.[36] If DNA repair is deficient, DNA damage tends to accumulate. Such excess DNA damage may increase mutational errors during DNA replication due to error-prone translesion synthesis. Excess DNA damage may also increase epigenetic alterations due to errors during DNA repair.[37][38] Such mutations and epigenetic alterations may give rise to cancer. The frequent epigenetic deficiency of ATM in a number of cancers likely contributed to the progression of those cancers.
Meiosis
ATM functions during meiotic prophase.[39] The wild-type ATM gene is expressed at a four-fold increased level in human testes compared to somatic cells (such as skin fibroblasts).[40] In both mice and humans, ATM deficiency results in female and male infertility. Deficient ATM expression causes severe meiotic disruption during prophase I.[41] In addition, impaired ATM-mediated DNA DSB repair has been identified as a likely cause of aging of mouse and human oocytes.[42] Expression of the ATM gene, as well as other key DSB repair genes, declines with age in mouse and human oocytes and this decline is paralleled by an increase of DSBs in primordial follicles.[42] These findings indicate that ATM-mediated homologous recombinational repair is a crucial function of meiosis.
Inhibitors
Several ATM kinase inhibitors are currently known, some of which are already in clinical trials.[43][44][45] One of the first discovered ATM inhibitors is caffeine with an IC50 of 0.2 mM and only a low selectivity within the PIKK family.[46][47] Wortmannin is an irreversible inhibitor of ATM with no selectivity over other related PIKK and PI3K kinases.[48] The most important group of inhibitors are compounds based on the 3-methyl-1,3-dihydro-2H-imidazo[4,5-c]quinolin-2-one scaffold. The first important representative is the inhibitor is Dactolisib (NVP-BEZ235), which was first published by Novartis as a selective mTOR/PI3K inhibitor.[49] It was later shown to also inhibit other PIKK kinases such as ATM, DNA-PK and ATR.[50] Various optimisation efforts by AstraZeneca (AZD0156, AZD1390), Merck (M4076) and Dimitrov et al. have led to highly active ATM inhibitors with greater potency.[51][52][53]


Interactions
Ataxia telangiectasia mutated has been shown to interact with:
Tefu
The Tefu protein of Drosophila melanogaster is a structural and functional homolog of the human ATM protein.[78] Tefu, like ATM, is required for DNA repair and normal levels of meiotic recombination in oocytes.
See also
References
- 1 2 3 GRCh38: Ensembl release 89: ENSG00000149311 - Ensembl, May 2017
- 1 2 3 GRCm38: Ensembl release 89: ENSMUSG00000034218 - Ensembl, May 2017
- ↑ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ↑ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ↑ Lee, Ji-Hoon; Paull, Tanya T. (December 2021). "Cellular functions of the protein kinase ATM and their relevance to human disease". Nature Reviews Molecular Cell Biology. 22 (12): 796–814. doi:10.1038/s41580-021-00394-2. ISSN 1471-0080. PMID 34429537. S2CID 237294441.
- ↑ Savitsky K, Bar-Shira A, Gilad S, Rotman G, Ziv Y, Vanagaite L, et al. (June 1995). "A single ataxia telangiectasia gene with a product similar to PI-3 kinase". Science. 268 (5218): 1749–53. Bibcode:1995Sci...268.1749S. doi:10.1126/science.7792600. PMID 7792600.
- ↑ "Entrez Gene: ATM ataxia telangiectasia mutated (includes complementation groups A, C and D)".
- 1 2 Banin S, Moyal L, Shieh S, Taya Y, Anderson CW, Chessa L, et al. (September 1998). "Enhanced phosphorylation of p53 by ATM in response to DNA damage". Science. 281 (5383): 1674–7. Bibcode:1998Sci...281.1674B. doi:10.1126/science.281.5383.1674. PMID 9733514.
- 1 2 Canman CE, Lim DS, Cimprich KA, Taya Y, Tamai K, Sakaguchi K, et al. (September 1998). "Activation of the ATM kinase by ionizing radiation and phosphorylation of p53". Science. 281 (5383): 1677–9. Bibcode:1998Sci...281.1677C. doi:10.1126/science.281.5383.1677. PMID 9733515.
- 1 2 Lee JH, Paull TT (December 2007). "Activation and regulation of ATM kinase activity in response to DNA double-strand breaks". Oncogene. 26 (56): 7741–8. doi:10.1038/sj.onc.1210872. PMID 18066086.
- ↑ "Serine-protein kinase ATM - Homo sapiens (Human)".
- ↑ Stakyte, K.; Rotheneder, M.; Lammens, K.; Bartho, J. D.; Grädler, U.; Fuchß, T.; Pehl, U.; Alt, A.; van de Logt, E.; Hopfner, K. P. (October 2021). "Molecular basis of human ATM kinase inhibition". Nature Structural & Molecular Biology. 28 (10): 789–798. doi:10.1038/s41594-021-00654-x. ISSN 1545-9985. PMID 34556870. S2CID 237615473.
- 1 2 3 Lempiäinen H, Halazonetis TD (October 2009). "Emerging common themes in regulation of PIKKs and PI3Ks". The EMBO Journal. 28 (20): 3067–73. doi:10.1038/emboj.2009.281. PMC 2752028. PMID 19779456.
- ↑ Huang X, Halicka HD, Darzynkiewicz Z (November 2004). "Detection of histone H2AX phosphorylation on Ser-139 as an indicator of DNA damage (DNA double-strand breaks)". Current Protocols in Cytometry. Chapter 7: Unit 7.27. doi:10.1002/0471142956.cy0727s30. ISBN 0-471-14295-6. PMID 18770804. S2CID 32281084.
- 1 2 Morgan DO (2007). The cell cycle: Principles of Control. Oxford University Press. ISBN 978-0-19-920610-0.
- ↑ Valentin-Vega YA, Maclean KH, Tait-Mulder J, Milasta S, Steeves M, Dorsey FC, et al. (February 2012). "Mitochondrial dysfunction in ataxia-telangiectasia". Blood. 119 (6): 1490–500. doi:10.1182/blood-2011-08-373639. PMC 3286212. PMID 22144182.
- ↑ Datan E, Roy SG, Germain G, Zali N, McLean JE, Golshan G, et al. (March 2016). "Dengue-induced autophagy, virus replication and protection from cell death require ER stress (PERK) pathway activation". Cell Death & Disease. 7 (e2127): e2127. doi:10.1038/cddis.2015.409. PMC 4823927. PMID 26938301.
- ↑ Bakkenist CJ, Kastan MB (January 2003). "DNA damage activates ATM through intermolecular autophosphorylation and dimer dissociation". Nature. 421 (6922): 499–506. Bibcode:2003Natur.421..499B. doi:10.1038/nature01368. PMID 12556884. S2CID 4403303.
- ↑ Hall MJ, Bernhisel R, Hughes E, Larson K, Rosenthal ET, Singh NA, et al. (April 2021). "Germline Pathogenic Variants in the Ataxia Telangiectasia Mutated (ATM) Gene are Associated with High and Moderate Risks for Multiple Cancers". Cancer Prevention Research. 14 (4): 433–440. doi:10.1158/1940-6207.CAPR-20-0448. PMC 8026745. PMID 33509806.
- ↑ Canman CE, Lim DS (December 1998). "The role of ATM in DNA damage responses and cancer". Oncogene. 17 (25): 3301–8. doi:10.1038/sj.onc.1202577. PMID 9916992.
- ↑ Kurz EU, Lees-Miller SP (2004). "DNA damage-induced activation of ATM and ATM-dependent signaling pathways". DNA Repair. 3 (8–9): 889–900. doi:10.1016/j.dnarep.2004.03.029. PMID 15279774.
- 1 2 Chen J (September 2000). "Ataxia telangiectasia-related protein is involved in the phosphorylation of BRCA1 following deoxyribonucleic acid damage". Cancer Research. 60 (18): 5037–9. PMID 11016625.
- ↑ Cremona CA, Behrens A (June 2014). "ATM signalling and cancer". Oncogene. 33 (26): 3351–60. doi:10.1038/onc.2013.275. PMID 23851492.
- ↑ Friedenson B (August 2007). "The BRCA1/2 pathway prevents hematologic cancers in addition to breast and ovarian cancers". BMC Cancer. 7: 152. doi:10.1186/1471-2407-7-152. PMC 1959234. PMID 17683622.
- ↑ Armstrong SA, Schultz CW, Azimi-Sadjadi A, Brody JR, Pishvaian MJ (November 2019). "ATM Dysfunction in Pancreatic Adenocarcinoma and Associated Therapeutic Implications". Molecular Cancer Therapeutics. 18 (11): 1899–1908. doi:10.1158/1535-7163.MCT-19-0208. PMC 6830515. PMID 31676541.
- ↑ Dunlop CR, Wallez Y, Johnson TI, Bernaldo de Quirós Fernández S, Durant ST, Cadogan EB, et al. (August 2020). "Complete loss of ATM function augments replication catastrophe induced by ATR inhibition and gemcitabine in pancreatic cancer models". British Journal of Cancer. 123 (9): 1424–1436. doi:10.1038/s41416-020-1016-2. PMC 7591912. PMID 32741974. S2CID 220931196.
- ↑ Kwok M, Davies N, Agathanggelou A, Smith E, Oldreive C, Petermann E, et al. (February 2016). "ATR inhibition induces synthetic lethality and overcomes chemoresistance in TP53- or ATM-defective chronic lymphocytic leukemia cells". Blood. 127 (5): 582–95. doi:10.1182/blood-2015-05-644872. PMID 26563132.
- ↑ Min A, Im SA, Jang H, Kim S, Lee M, Kim DK, et al. (April 2017). "AZD6738, A Novel Oral Inhibitor of ATR, Induces Synthetic Lethality with ATM Deficiency in Gastric Cancer Cells". Molecular Cancer Therapeutics. 16 (4): 566–577. doi:10.1158/1535-7163.MCT-16-0378. PMID 28138034. S2CID 34149115.
- ↑ Vendetti FP, Lau A, Schamus S, Conrads TP, O'Connor MJ, Bakkenist CJ (December 2015). "The orally active and bioavailable ATR kinase inhibitor AZD6738 potentiates the anti-tumor effects of cisplatin to resolve ATM-deficient non-small cell lung cancer in vivo". Oncotarget. 6 (42): 44289–305. doi:10.18632/oncotarget.6247. PMC 4792557. PMID 26517239.
- ↑ Mehdipour P, Karami F, Javan F, Mehrazin M (August 2015). "Linking ATM Promoter Methylation to Cell Cycle Protein Expression in Brain Tumor Patients: Cellular Molecular Triangle Correlation in ATM Territory". Molecular Neurobiology. 52 (1): 293–302. doi:10.1007/s12035-014-8864-9. PMID 25159481. S2CID 35412479.
- ↑ Delmonico L, dos Santos Moreira A, Franco MF, Esteves EB, Scherrer L, Gallo CV, et al. (October 2015). "CDKN2A (p14(ARF)/p16(INK4a)) and ATM promoter methylation in patients with impalpable breast lesions". Human Pathology. 46 (10): 1540–7. doi:10.1016/j.humpath.2015.06.016. PMID 26255234.
- ↑ Vo QN, Kim WJ, Cvitanovic L, Boudreau DA, Ginzinger DG, Brown KD (December 2004). "The ATM gene is a target for epigenetic silencing in locally advanced breast cancer". Oncogene. 23 (58): 9432–7. doi:10.1038/sj.onc.1208092. PMID 15516988.
- ↑ Safar AM, Spencer H, Su X, Coffey M, Cooney CA, Ratnasinghe LD, et al. (June 2005). "Methylation profiling of archived non-small cell lung cancer: a promising prognostic system". Clinical Cancer Research. 11 (12): 4400–5. doi:10.1158/1078-0432.CCR-04-2378. PMID 15958624.
- ↑ Bolt J, Vo QN, Kim WJ, McWhorter AJ, Thomson J, Hagensee ME, et al. (November 2005). "The ATM/p53 pathway is commonly targeted for inactivation in squamous cell carcinoma of the head and neck (SCCHN) by multiple molecular mechanisms". Oral Oncology. 41 (10): 1013–20. doi:10.1016/j.oraloncology.2005.06.003. PMID 16139561.
- ↑ Kastan MB (April 2008). "DNA damage responses: mechanisms and roles in human disease: 2007 G.H.A. Clowes Memorial Award Lecture". Molecular Cancer Research. 6 (4): 517–24. doi:10.1158/1541-7786.MCR-08-0020. PMID 18403632.
- ↑ Harper JW, Elledge SJ (December 2007). "The DNA damage response: ten years after". Molecular Cell. 28 (5): 739–45. doi:10.1016/j.molcel.2007.11.015. PMID 18082599.
- ↑ O'Hagan HM, Mohammad HP, Baylin SB (August 2008). "Double strand breaks can initiate gene silencing and SIRT1-dependent onset of DNA methylation in an exogenous promoter CpG island". PLOS Genetics. 4 (8): e1000155. doi:10.1371/journal.pgen.1000155. PMC 2491723. PMID 18704159.
- ↑ Cuozzo C, Porcellini A, Angrisano T, Morano A, Lee B, Di Pardo A, et al. (July 2007). "DNA damage, homology-directed repair, and DNA methylation". PLOS Genetics. 3 (7): e110. doi:10.1371/journal.pgen.0030110. PMC 1913100. PMID 17616978.
- ↑ Hamer G, Kal HB, Westphal CH, Ashley T, de Rooij DG (April 2004). "Ataxia telangiectasia mutated expression and activation in the testis". Biology of Reproduction. 70 (4): 1206–12. doi:10.1095/biolreprod.103.024950. PMID 14681204.
- ↑ Galetzka D, Weis E, Kohlschmidt N, Bitz O, Stein R, Haaf T (April 2007). "Expression of somatic DNA repair genes in human testes". Journal of Cellular Biochemistry. 100 (5): 1232–9. doi:10.1002/jcb.21113. PMID 17177185. S2CID 23743474.
- ↑ Barlow C, Liyanage M, Moens PB, Tarsounas M, Nagashima K, Brown K, et al. (October 1998). "Atm deficiency results in severe meiotic disruption as early as leptonema of prophase I". Development. 125 (20): 4007–17. doi:10.1242/dev.125.20.4007. PMID 9735362.
- 1 2 Titus S, Li F, Stobezki R, Akula K, Unsal E, Jeong K, et al. (February 2013). "Impairment of BRCA1-related DNA double-strand break repair leads to ovarian aging in mice and humans". Science Translational Medicine. 5 (172): 172ra21. doi:10.1126/scitranslmed.3004925. PMC 5130338. PMID 23408054.
- ↑ "CTG Labs - NCBI". clinicaltrials.gov. Retrieved 2023-08-29.
- ↑ "CTG Labs - NCBI". clinicaltrials.gov. Retrieved 2023-08-29.
- ↑ "CTG Labs - NCBI". clinicaltrials.gov. Retrieved 2023-08-29.
- ↑ Blasina A, Price BD, Turenne GA, McGowan CH (October 1999). "Caffeine inhibits the checkpoint kinase ATM". Current Biology. 9 (19): 1135–1138. doi:10.1016/s0960-9822(99)80486-2. PMID 10531013.
- ↑ Sarkaria JN, Busby EC, Tibbetts RS, Roos P, Taya Y, Karnitz LM, Abraham RT (September 1999). "Inhibition of ATM and ATR kinase activities by the radiosensitizing agent, caffeine". Cancer Research. 59 (17): 4375–4382. PMID 10485486.
- ↑ Sarkaria JN, Tibbetts RS, Busby EC, Kennedy AP, Hill DE, Abraham RT (October 1998). "Inhibition of phosphoinositide 3-kinase related kinases by the radiosensitizing agent wortmannin". Cancer Research. 58 (19): 4375–4382. PMID 9766667.
- ↑ Maira SM, Stauffer F, Brueggen J, Furet P, Schnell C, Fritsch C, et al. (July 2008). "Identification and characterization of NVP-BEZ235, a new orally available dual phosphatidylinositol 3-kinase/mammalian target of rapamycin inhibitor with potent in vivo antitumor activity". Molecular Cancer Therapeutics. 7 (7): 1851–1863. doi:10.1158/1535-7163.MCT-08-0017. PMID 18606717.
- ↑ Toledo LI, Murga M, Zur R, Soria R, Rodriguez A, Martinez S, et al. (June 2011). "A cell-based screen identifies ATR inhibitors with synthetic lethal properties for cancer-associated mutations". Nature Structural & Molecular Biology. 18 (6): 721–727. doi:10.1038/nsmb.2076. PMC 4869831. PMID 21552262.
- ↑ Pike KG, Barlaam B, Cadogan E, Campbell A, Chen Y, Colclough N, et al. (May 2018). "The Identification of Potent, Selective, and Orally Available Inhibitors of Ataxia Telangiectasia Mutated (ATM) Kinase: The Discovery of AZD0156 (8-{6-[3-(Dimethylamino)propoxy]pyridin-3-yl}-3-methyl-1-(tetrahydro-2 H-pyran-4-yl)-1,3-dihydro-2 H-imidazo[4,5- c]quinolin-2-one)". Journal of Medicinal Chemistry. 61 (9): 3823–3841. doi:10.1021/acs.jmedchem.7b01896. PMID 29683659.
- ↑ Zimmermann A, Zenke FT, Chiu LY, Dahmen H, Pehl U, Fuchss T, et al. (June 2022). "A New Class of Selective ATM Inhibitors as Combination Partners of DNA Double-Strand Break Inducing Cancer Therapies". Molecular Cancer Therapeutics. 21 (6): 859–870. doi:10.1158/1535-7163.MCT-21-0934. PMC 9381122. PMID 35405736.
- ↑ Dimitrov T, Anli C, Moschopoulou AA, Kronenberger T, Kudolo M, Geibel C, et al. (May 2022). "Development of novel urea-based ATM kinase inhibitors with subnanomolar cellular potency and high kinome selectivity". European Journal of Medicinal Chemistry. 235: 114234. doi:10.1016/j.ejmech.2022.114234. PMID 35325634. S2CID 247356817.
- 1 2 Chen G, Yuan SS, Liu W, Xu Y, Trujillo K, Song B, et al. (April 1999). "Radiation-induced assembly of Rad51 and Rad52 recombination complex requires ATM and c-Abl". The Journal of Biological Chemistry. 274 (18): 12748–52. doi:10.1074/jbc.274.18.12748. PMID 10212258.
- 1 2 Kishi S, Zhou XZ, Ziv Y, Khoo C, Hill DE, Shiloh Y, Lu KP (August 2001). "Telomeric protein Pin2/TRF1 as an important ATM target in response to double strand DNA breaks". The Journal of Biological Chemistry. 276 (31): 29282–91. doi:10.1074/jbc.M011534200. PMID 11375976.
- ↑ Shafman T, Khanna KK, Kedar P, Spring K, Kozlov S, Yen T, et al. (May 1997). "Interaction between ATM protein and c-Abl in response to DNA damage". Nature. 387 (6632): 520–3. Bibcode:1997Natur.387R.520S. doi:10.1038/387520a0. PMID 9168117. S2CID 4334242.
- 1 2 3 4 5 6 7 Kim ST, Lim DS, Canman CE, Kastan MB (December 1999). "Substrate specificities and identification of putative substrates of ATM kinase family members". The Journal of Biological Chemistry. 274 (53): 37538–43. doi:10.1074/jbc.274.53.37538. PMID 10608806.
- 1 2 3 4 Wang Y, Cortez D, Yazdi P, Neff N, Elledge SJ, Qin J (April 2000). "BASC, a super complex of BRCA1-associated proteins involved in the recognition and repair of aberrant DNA structures". Genes & Development. 14 (8): 927–39. doi:10.1101/gad.14.8.927. PMC 316544. PMID 10783165.
- ↑ Gatei M, Scott SP, Filippovitch I, Soronika N, Lavin MF, Weber B, Khanna KK (June 2000). "Role for ATM in DNA damage-induced phosphorylation of BRCA1". Cancer Research. 60 (12): 3299–304. PMID 10866324.
- ↑ Cortez D, Wang Y, Qin J, Elledge SJ (November 1999). "Requirement of ATM-dependent phosphorylation of brca1 in the DNA damage response to double-strand breaks". Science. 286 (5442): 1162–6. doi:10.1126/science.286.5442.1162. PMID 10550055.
- ↑ Tibbetts RS, Cortez D, Brumbaugh KM, Scully R, Livingston D, Elledge SJ, Abraham RT (December 2000). "Functional interactions between BRCA1 and the checkpoint kinase ATR during genotoxic stress". Genes & Development. 14 (23): 2989–3002. doi:10.1101/gad.851000. PMC 317107. PMID 11114888.
- ↑ Gatei M, Zhou BB, Hobson K, Scott S, Young D, Khanna KK (May 2001). "Ataxia telangiectasia mutated (ATM) kinase and ATM and Rad3 related kinase mediate phosphorylation of Brca1 at distinct and overlapping sites. In vivo assessment using phospho-specific antibodies" (PDF). The Journal of Biological Chemistry. 276 (20): 17276–80. doi:10.1074/jbc.M011681200. PMID 11278964. S2CID 43554268.
- ↑ Beamish H, Kedar P, Kaneko H, Chen P, Fukao T, Peng C, et al. (August 2002). "Functional link between BLM defective in Bloom's syndrome and the ataxia-telangiectasia-mutated protein, ATM". The Journal of Biological Chemistry. 277 (34): 30515–23. doi:10.1074/jbc.M203801200. PMID 12034743.
- ↑ Suzuki K, Kodama S, Watanabe M (September 1999). "Recruitment of ATM protein to double strand DNA irradiated with ionizing radiation". The Journal of Biological Chemistry. 274 (36): 25571–5. doi:10.1074/jbc.274.36.25571. PMID 10464290.
- ↑ Taniguchi T, Garcia-Higuera I, Xu B, Andreassen PR, Gregory RC, Kim ST, et al. (May 2002). "Convergence of the fanconi anemia and ataxia telangiectasia signaling pathways". Cell. 109 (4): 459–72. doi:10.1016/s0092-8674(02)00747-x. PMID 12086603. S2CID 16580666.
- ↑ Reuter TY, Medhurst AL, Waisfisz Q, Zhi Y, Herterich S, Hoehn H, et al. (October 2003). "Yeast two-hybrid screens imply involvement of Fanconi anemia proteins in transcription regulation, cell signaling, oxidative metabolism, and cellular transport". Experimental Cell Research. 289 (2): 211–21. doi:10.1016/s0014-4827(03)00261-1. PMID 14499622.
- ↑ Kang J, Ferguson D, Song H, Bassing C, Eckersdorff M, Alt FW, Xu Y (January 2005). "Functional interaction of H2AX, NBS1, and p53 in ATM-dependent DNA damage responses and tumor suppression". Molecular and Cellular Biology. 25 (2): 661–70. doi:10.1128/MCB.25.2.661-670.2005. PMC 543410. PMID 15632067.
- ↑ Fabbro M, Savage K, Hobson K, Deans AJ, Powell SN, McArthur GA, Khanna KK (July 2004). "BRCA1-BARD1 complexes are required for p53Ser-15 phosphorylation and a G1/S arrest following ionizing radiation-induced DNA damage". The Journal of Biological Chemistry. 279 (30): 31251–8. doi:10.1074/jbc.M405372200. PMID 15159397.
- ↑ Khanna KK, Keating KE, Kozlov S, Scott S, Gatei M, Hobson K, et al. (December 1998). "ATM associates with and phosphorylates p53: mapping the region of interaction". Nature Genetics. 20 (4): 398–400. doi:10.1038/3882. PMID 9843217. S2CID 23994762.
- ↑ Westphal CH, Schmaltz C, Rowan S, Elson A, Fisher DE, Leder P (May 1997). "Genetic interactions between atm and p53 influence cellular proliferation and irradiation-induced cell cycle checkpoints". Cancer Research. 57 (9): 1664–7. PMID 9135004.
- ↑ Bao S, Tibbetts RS, Brumbaugh KM, Fang Y, Richardson DA, Ali A, et al. (June 2001). "ATR/ATM-mediated phosphorylation of human Rad17 is required for genotoxic stress responses". Nature. 411 (6840): 969–74. Bibcode:2001Natur.411..969B. doi:10.1038/35082110. PMID 11418864. S2CID 4429058.
- ↑ Li S, Ting NS, Zheng L, Chen PL, Ziv Y, Shiloh Y, et al. (July 2000). "Functional link of BRCA1 and ataxia telangiectasia gene product in DNA damage response". Nature. 406 (6792): 210–5. Bibcode:2000Natur.406..210L. doi:10.1038/35018134. PMID 10910365. S2CID 3266654.
- ↑ Long X, Lin Y, Ortiz-Vega S, Yonezawa K, Avruch J (April 2005). "Rheb binds and regulates the mTOR kinase". Current Biology. 15 (8): 702–13. doi:10.1016/j.cub.2005.02.053. PMID 15854902. S2CID 3078706.
- ↑ Chang L, Zhou B, Hu S, Guo R, Liu X, Jones SN, Yen Y (November 2008). "ATM-mediated serine 72 phosphorylation stabilizes ribonucleotide reductase small subunit p53R2 protein against MDM2 to DNA damage". Proceedings of the National Academy of Sciences of the United States of America. 105 (47): 18519–24. Bibcode:2008PNAS..10518519C. doi:10.1073/pnas.0803313105. PMC 2587585. PMID 19015526.
- ↑ Kim ST, Xu B, Kastan MB (March 2002). "Involvement of the cohesin protein, Smc1, in Atm-dependent and independent responses to DNA damage". Genes & Development. 16 (5): 560–70. doi:10.1101/gad.970602. PMC 155347. PMID 11877376.
- ↑ Fernandez-Capetillo O, Chen HT, Celeste A, Ward I, Romanienko PJ, Morales JC, et al. (December 2002). "DNA damage-induced G2-M checkpoint activation by histone H2AX and 53BP1". Nature Cell Biology. 4 (12): 993–7. doi:10.1038/ncb884. PMID 12447390. S2CID 12380387.
- ↑ Ward IM, Minn K, Jorda KG, Chen J (May 2003). "Accumulation of checkpoint protein 53BP1 at DNA breaks involves its binding to phosphorylated histone H2AX". The Journal of Biological Chemistry. 278 (22): 19579–82. doi:10.1074/jbc.C300117200. PMID 12697768.
- ↑ Oikemus SR, McGinnis N, Queiroz-Machado J, Tukachinsky H, Takada S, Sunkel CE, Brodsky MH (August 2004). "Drosophila atm/telomere fusion is required for telomeric localization of HP1 and telomere position effect". Genes & Development. 18 (15): 1850–61. doi:10.1101/gad.1202504. PMC 517405. PMID 15256487.
Further reading
- Giaccia AJ, Kastan MB (October 1998). "The complexity of p53 modulation: emerging patterns from divergent signals". Genes & Development. 12 (19): 2973–83. doi:10.1101/gad.12.19.2973. PMID 9765199.
- Akst J (2015). "Another Telomere-Regulating Enzyme Found". The Scientist (November 12).
- Kastan MB, Lim DS (December 2000). "The many substrates and functions of ATM". Nature Reviews. Molecular Cell Biology. 1 (3): 179–86. doi:10.1038/35043058. PMID 11252893. S2CID 10691352.
- Shiloh Y (2002). "ATM: From Phenotype to Functional Genomics — and Back". The Human Genome. pp. 51–70. doi:10.1007/978-3-662-04667-8_4. ISBN 978-3-662-04669-2. PMID 11859564.
{{cite book}}:|journal=ignored (help) - Redon C, Pilch D, Rogakou E, Sedelnikova O, Newrock K, Bonner W (April 2002). "Histone H2A variants H2AX and H2AZ". Current Opinion in Genetics & Development. 12 (2): 162–9. doi:10.1016/S0959-437X(02)00282-4. PMID 11893489.
- Tang Y (February 2002). "[ATM and Cancer]". Zhongguo Shi Yan Xue Ye Xue Za Zhi. 10 (1): 77–80. PMID 12513844.
- Shiloh Y (March 2003). "ATM and related protein kinases: safeguarding genome integrity". Nature Reviews. Cancer. 3 (3): 155–68. doi:10.1038/nrc1011. PMID 12612651. S2CID 22770833.
- Gumy-Pause F, Wacker P, Sappino AP (February 2004). "ATM gene and lymphoid malignancies". Leukemia. 18 (2): 238–42. doi:10.1038/sj.leu.2403221. PMID 14628072.
- Kurz EU, Lees-Miller SP (2005). "DNA damage-induced activation of ATM and ATM-dependent signaling pathways". DNA Repair. 3 (8–9): 889–900. doi:10.1016/j.dnarep.2004.03.029. PMID 15279774.
- Abraham RT (2005). "The ATM-related kinase, hSMG-1, bridges genome and RNA surveillance pathways". DNA Repair. 3 (8–9): 919–25. doi:10.1016/j.dnarep.2004.04.003. PMID 15279777.
- Lavin MF, Scott S, Gueven N, Kozlov S, Peng C, Chen P (2005). "Functional consequences of sequence alterations in the ATM gene". DNA Repair. 3 (8–9): 1197–205. doi:10.1016/j.dnarep.2004.03.011. PMID 15279808.
- Meulmeester E, Pereg Y, Shiloh Y, Jochemsen AG (September 2005). "ATM-mediated phosphorylations inhibit Mdmx/Mdm2 stabilization by HAUSP in favor of p53 activation". Cell Cycle. 4 (9): 1166–70. doi:10.4161/cc.4.9.1981. PMID 16082221.
- Ahmed M, Rahman N (September 2006). "ATM and breast cancer susceptibility". Oncogene. 25 (43): 5906–11. doi:10.1038/sj.onc.1209873. PMID 16998505.
External links
- https://web.archive.org/web/20060107000211/http://www.hprd.org/protein/06347
- Drosophila telomere fusion - The Interactive Fly
- GeneReviews/NCBI/NIH/UW entry on Ataxia telangiectasia
- OMIM entries on Ataxia telangiectasia
- Human ATM genome location and ATM gene details page in the UCSC Genome Browser.
- Overview of all the structural information available in the PDB for UniProt: Q13315 (Serine-protein kinase ATM) at the PDBe-KB.