Hypersensitive response (HR) is a mechanism used by plants to prevent the spread of infection by microbial pathogens. HR is characterized by the rapid death of cells in the local region surrounding an infection and it serves to restrict the growth and spread of pathogens to other parts of the plant. It is analogous to the innate immune system found in animals, and commonly precedes a slower systemic (whole plant) response, which ultimately leads to systemic acquired resistance (SAR).[1] HR can be observed in the vast majority of plant species and is induced by a wide range of plant pathogens such as oomycetes, viruses, fungi and even insects.[2]

HR is commonly thought of as an effective defence strategy against biotrophic plant pathogens, which require living tissue to gain nutrients. In the case of necrotrophic pathogens, HR might even be beneficial to the pathogen, as they require dead plant cells to obtain nutrients. The situation becomes complicated when considering pathogens such as Phytophthora infestans which at the initial stages of the infection act as biotrophs but later switch to a necrotrophic lifestyle. It is proposed that in this case HR might be beneficial in the early stages of the infection but not in the later stages.[3]
Genetics
The first idea of how the hypersensitive response occurs came from Harold Henry Flor's gene-for-gene model. He postulated that for every resistance (R) gene encoded by the plant, there is a corresponding avirulence (Avr) gene encoded by the microbe. The plant is resistant to the pathogen if both the Avr and R genes are present during the plant-pathogen interaction.[4] The genes that are involved in the plant-pathogen interactions tend to evolve at a very rapid rate.[5]
Very often, the resistance mediated by R genes is due to them inducing HR, which leads to apoptosis. Most plant R genes encode NOD-like receptor (NLR) proteins.[6] NLR protein domain architecture consists of an NB-ARC domain which is a nucleotide-binding domain, responsible for conformational changes associated with the activation of the NLR protein. In the inactive form, the NB-ARC domain is bound to Adenosine diphosphate (ADP). When a pathogen is sensed, the ADP is exchanged for Adenosine triphosphate (ATP) and this induces a conformational change in the NLR protein, which results in HR. At the N-terminus, the NLR either has a Toll-Interleukin receptor (TIR) domain (also found in mammalian toll-like receptors) or a coiled-coil (CC) motif. Both TIR and CC domains are implicated in causing cell death during HR. The C-terminus of the NLRs consists of a leucine-rich repeat (LRR) motif, which is involved in sensing the pathogen virulence factors.[7]
Mechanism
HR is triggered by the plant when it recognizes a pathogen. The identification of a pathogen typically occurs when a virulence gene product, secreted by a pathogen, binds to, or indirectly interacts with the product of a plant R gene. R genes are highly polymorphic, and many plants produce several different types of R gene products, enabling them to recognize virulence products produced by many different pathogens.[8]
In phase one of the HR, the activation of R genes triggers an ion flux, involving an efflux of hydroxide and potassium to the outside the cells, and an influx of calcium and hydrogen ions into the cells.[9]
In phase two, the cells involved in the HR generate an oxidative burst by producing reactive oxygen species (ROS), superoxide anions, hydrogen peroxide, hydroxyl radicals and nitrous oxide. These compounds affect cellular membrane function, in part by inducing lipid peroxidation and by causing lipid damage.[9]
The alteration of ion components in the cell and the breakdown of cellular components in the presence of ROS result in the death of affected cells, as well as the formation of local lesions. Reactive oxygen species also trigger the deposition of lignin and callose, as well as the cross-linking of pre-formed hydroxyproline-rich glycoproteins such as P33 to the wall matrix via the tyrosine in the PPPPY motif.[9] These compounds serve to reinforce the walls of cells surrounding the infection, creating a barrier and inhibiting the spread of the infection.[10] Activation of HR also results in disruption of the cytoskeleton, mitochondrial function and metabolic changes, all of which might be implicated in causing cell death.[11][12][13]
Direct and indirect activation
HR can be activated in two main ways: directly and indirectly. Direct binding of the virulence factors to the NLRs can result in the activation of HR. However, this seems to be quite rare. More commonly, the virulence factors target certain cellular proteins that they modify and this modification is then sensed by NLRs. Indirect recognition seems to be more common as multiple virulence factors can modify the same cellular protein with the same modifications thus allowing one receptor to recognize multiple virulence factors.[14] Sometimes, the protein domains targeted by the virulence factors are integrated into the NLRs. An example of this can be observed in plant resistance to the rice blast pathogen, where the RGA5 NLR has a heavy-metal-associated (HMA) domain integrated into its structure, which is targeted by multiple effector proteins.[15]
An example of indirect recognition: AvrPphB is a type III effector protein secreted by Pseudomonas syringae. This is a protease which cleaves a cellular kinase called PBS1. The modified kinase is sensed by RPS5 NLR.[16]
The Resistosome
Recent structural studies of CC-NLR proteins have suggested that after the virulence factors are sensed, the NLRs assemble into a pentameric structure known as the resistosome. The resistosome seems to have a high affinity for the cellular membrane. When the resistosome is assembled, a helix sticks out from the N-terminus of each NLR and this creates a pore in the membrane which allows leakage of ions to occur and thus the cell dies. However, this mechanism is only inferred from the structure and there are currently no mechanistic studies to support this. It is still not known how the TIR-NLR proteins are activated. Recent research suggests that they require CC-NLR proteins downstream of them, which are then activated to form the resistosomes and induce HR.[17]
NLR pairs and networks
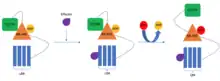
It is known that NLRs can function individually but there are also cases where the NLR proteins work in pairs. The pair consists of a sensor NLR and a helper NLR. The sensor NLR is responsible for recognizing the pathogen secreted effector protein and activating the helper NLR which then executes the cell death. The genes of both the sensor and the respective helper NLR are usually paired in the genome and their expression could be controlled by the same promoter. This allows the functional pair, instead of individual components, to be segregated during cell division and also ensures that equal amounts of both NLRs are made in the cell.[18]
The receptor pairs work through two main mechanisms: negative regulation or cooperation.
In the negative regulation scenario, the sensor NLR is responsible for negatively regulating the helper NLR and preventing cell death under normal conditions. However, when the effector protein is introduced and recognized by the sensor NLR, the negative regulation of the helper NLR is relieved and HR is induced.[19]
In the cooperation mechanisms, when the sensor NLR recognizes the effector protein it signals to the helper NLR, thus activating it.[20]
Recently, it was discovered that in addition to acting as singletons or pairs, the plant NLRs can act in networks. In these networks, there are usually many sensor NLRs paired to relatively few helper NLRs.[20]
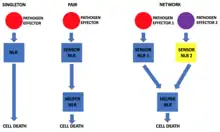
One example of proteins involved in NLR networks are those belonging to the NRC superclade. It seems that the networks evolved from a duplication event of a genetically linked NLR pair into an unlinked locus which allowed the new pair to evolve to respond to a new pathogen. This separation seems to provide plasticity to the system, as it allows the sensor NLRs to evolve more rapidly in response to the fast evolution of pathogen effectors whereas the helper NLR can evolve much slower to maintain its ability to induce HR. However, it seems that during evolution new helper NLRs also evolved, supposedly, because certain sensor NLRs require specific helper NLRs to function optimally.[20]
Bioinformatic analysis of plant NLRs has shown that there is a conserved MADA motif at the N-terminus of helper NLRs but not sensor NLRs. Around 20% of all CC-NLRs have the MADA motif, implying the motif's importance for the execution of HR.[21]
Regulation
Accidental activation of HR through the NLR proteins could cause vast destruction of the plant tissue, thus, the NLRs are kept in an inactive form through tight negative regulation at both transcriptional and post-translational levels. Under normal conditions, the mRNA of NLRs are transcribed at very low levels, which results in low levels of protein in the cell. The NLRs also require a considerable number of chaperone proteins for their folding. Misfolded proteins are immediately ubiquitinated and degraded by the proteasome.[22] It has been observed that in many cases, if the chaperone proteins involved in NLR biosynthesis are knocked-out, HR is abolished and NLR levels are significantly reduced.[23]
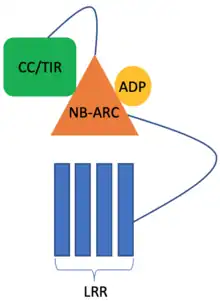
Intramolecular interactions are also essential for the regulation of HR. The NLR proteins are not linear: the NB-ARC domain is sandwiched in between the LRR and TIR/CC domains. Under normal conditions, there is a lot more ATP present in the cytoplasm than ADP, and this arrangement of the NLR proteins prevents the spontaneous exchange of ADP for ATP and thus activation of HR. Only when a virulence factor is sensed, the ADP is exchanged for ATP.[14]
Mutations in certain components of plant defence machinery result in HR being activated without the presence of pathogen effector proteins. Some of these mutations are observed in NLR genes and cause these NLR proteins to become auto-active due to disrupted intramolecular regulatory mechanisms. Other mutations causing spontaneous HR are present in proteins involved in ROS production during pathogen invasion.[3]
HR is also a temperature-sensitive process and it has been observed that in many cases plant-pathogen interactions do not induce HR at temperatures above 30 °C, which subsequently leads to increased susceptibility to the pathogen.[24] The mechanisms behind the influence of temperature on plant resistance to pathogens are not understood in detail, however, research suggests that the NLR protein levels might be important in this regulation.[25] It is also proposed that at higher temperatures the NLR proteins are less likely to form oligomeric complexes, thus inhibiting their ability to induce HR.[26]
It has also been shown that HR is dependent on the light conditions, which could be linked to the activity of chloroplasts and mainly their ability to generate ROS.[27]
Mediators
Several enzymes have been shown to be involved in generation of ROS. For example, copper amine oxidase, catalyzes the oxidative deamination of polyamines, especially putrescine, and releases the ROS mediators hydrogen peroxide and ammonia.[28] Other enzymes thought to play a role in ROS production include xanthine oxidase, NADPH oxidase, oxalate oxidase, peroxidases, and flavin containing amine oxidases.[9]
In some cases, the cells surrounding the lesion synthesize antimicrobial compounds, including phenolics, phytoalexins, and pathogenesis related (PR) proteins, including β-glucanases and chitinases. These compounds may act by puncturing bacterial cell walls; or by delaying maturation, disrupting metabolism, or preventing reproduction of the pathogen in question.
Studies have suggested that the actual mode and sequence of the dismantling of plant cellular components depends on each individual plant-pathogen interaction, but all HR seem to require the involvement of cysteine proteases. The induction of cell death and the clearance of pathogens also requires active protein synthesis, an intact actin cytoskeleton, and the presence of salicylic acid.[8]
Pathogen evasion
Pathogens have evolved several strategies to suppress plant defense responses. Host processes usually targeted by bacteria include; alterations to programmed cell death pathways, inhibiting cell wall-based defenses, and altering plant hormone signaling and expression of defense genes.[29]
Systemic immunity
Local initiation of HR in response to certain necrotrophic pathogens has been shown to allow the plants to develop systemic immunity against the pathogen.[30] Scientists have been trying to exploit the ability of HR to induce systemic resistance in plants in order to create transgenic plants resistant to certain pathogens. Pathogen-inducible promoters have been linked to auto-active NLR genes to induce HR response only when the pathogen is present but not at any other time. This approach, however, has been mostly unfeasible as the modification also leads to a substantial reduction in plant yields.[3]
Hypersensitive response as a driver for plant speciation
It has been noticed in Arabidopsis that sometimes when two different plant lines are crossed together, the offspring show signs of hybrid necrosis. This is due to the parent plants containing incompatible NLRs, which when expressed together in the same cell, induce spontaneous HR.[31]
This observation raised a hypothesis that plant pathogens can lead to the speciation of plants – if plant populations from the same species develop incompatible NLRs in response to different pathogen effectors, this can lead to hybrid necrosis in the F1 offspring, which substantially reduces the fitness of the offspring and gene flow to subsequent generations.[32]
Comparison to animal innate immunity
Both plants and animals have NLR proteins which seem to have the same biological function – to induce cell death. The N-termini of plant and animal NLRs vary but it seems that both have LRR domains at the C-terminus.[33]
A big difference between animal and plant NLRs is in what they recognise. Animal NLRs mainly recognise pathogen-associated molecular patterns (PAMPs), while plant NLRs mostly recognise pathogen effector proteins. This makes sense as NLRs are present inside of the cell and plants rarely have intracellular pathogens, except for viruses and viruses do not have PAMPs as they are rapidly evolving. Animals, on the other hand, have intracellular pathogens.[34]
The vast majority of plant lineages, except for certain algae, such as Chlamydomonas, have NLRs. NLRs are also present in many animal species, however, they are not present in, for example, Drosophila melanogaster and Arthropods.[33]
Upon recognition of PAMPs by NLRs in animals, the NLRs oligomerise to form a structure known as the inflammasome, which activates pyroptosis. In plants, structural studies have suggested that the NLRs also oligomerise to form a structure called the resistosome, which also leads to cell death. It seems that in both plants and animals, the formation of the resistosome or the inflammasome, respectively, leads to cell death by forming pores in the membrane. It is inferred from protein structures that in plants the NLRs themselves are responsible for forming pores in the membrane, while in the case of the inflammasome, the pore-forming activity arises from gasdermin D which is cleaved by caspases as a result of the oligomerisation of the NLRs.[35][36] Plant cells do not have caspases.[37]
See also
References
- ↑ Freeman S (2003). "Chapter 37: Plant Defense Systems". Biological Science. Prentice Hall. Archived from the original on 2012-12-01. Retrieved 2007-01-12.
- ↑ Hammond-Kosack KE, Parker JE (April 2003). "Deciphering plant-pathogen communication: fresh perspectives for molecular resistance breeding". Current Opinion in Biotechnology. 14 (2): 177–93. doi:10.1016/S0958-1669(03)00035-1. PMID 12732319.
- 1 2 3 Balint-Kurti P (August 2019). "The plant hypersensitive response: concepts, control and consequences". Molecular Plant Pathology. 20 (8): 1163–1178. doi:10.1111/mpp.12821. PMC 6640183. PMID 31305008.
- ↑ Flor HH (September 1971). "Current Status of the Gene-For-Gene Concept". Annual Review of Phytopathology. 9 (1): 275–296. doi:10.1146/annurev.py.09.090171.001423. ISSN 0066-4286.
- ↑ Tiffin P, Moeller DA (December 2006). "Molecular evolution of plant immune system genes". Trends in Genetics. 22 (12): 662–70. doi:10.1016/j.tig.2006.09.011. PMID 17011664.
- ↑ Baggs E, Dagdas G, Krasileva KV (August 2017). "NLR diversity, helpers and integrated domains: making sense of the NLR IDentity". Current Opinion in Plant Biology. 38: 59–67. doi:10.1016/j.pbi.2017.04.012. PMID 28494248.
- ↑ Takken FL, Albrecht M, Tameling WI (August 2006). "Resistance proteins: molecular switches of plant defence". Current Opinion in Plant Biology. 9 (4): 383–90. doi:10.1016/j.pbi.2006.05.009. PMID 16713729.
- 1 2 Heath MC (October 2000). "Hypersensitive response-related death". Plant Molecular Biology. 44 (3): 321–34. doi:10.1023/A:1026592509060. PMID 11199391. S2CID 22107876.
- 1 2 3 4 Matthews B. "The Hypersensitive Response". Agricultural Research Service: Plant Science Institute. The United States Department of Agriculture. Archived from the original on 2007-02-22. Retrieved 2007-01-12.
- ↑ Pontier D, Balagué C, Roby D (September 1998). "The hypersensitive response. A programmed cell death associated with plant resistance". Comptes Rendus de l'Académie des Sciences, Série III. 321 (9): 721–34. Bibcode:1998CRASG.321..721P. doi:10.1016/s0764-4469(98)80013-9. PMID 9809204.
- ↑ Kobayashi I, Kobayashi Y, Hardham AR (December 1994). "Dynamic reorganization of microtubules and microfilaments in flax cells during the resistance response to flax rust infection". Planta. 195 (2). doi:10.1007/BF00199684. S2CID 36902627.
- ↑ Xie Z, Chen Z (February 2000). "Harpin-induced hypersensitive cell death is associated with altered mitochondrial functions in tobacco cells". Molecular Plant-Microbe Interactions. 13 (2): 183–90. doi:10.1094/MPMI.2000.13.2.183. PMID 10659708.
- ↑ Naton B, Hahlbrock K, Schmelzer E (September 1996). "Correlation of Rapid Cell Death with Metabolic Changes in Fungus-Infected, Cultured Parsley Cells". Plant Physiology. 112 (1): 433–444. doi:10.1104/pp.112.1.433. PMC 157965. PMID 12226400.
- 1 2 Bonardi V, Dangl JL (2012). "How complex are intracellular immune receptor signaling complexes?". Frontiers in Plant Science. 3: 237. doi:10.3389/fpls.2012.00237. PMC 3478704. PMID 23109935.
- ↑ Ortiz D, de Guillen K, Cesari S, Chalvon V, Gracy J, Padilla A, Kroj T (January 2017). "Magnaporthe oryzae Effector AVR-Pia by the Decoy Domain of the Rice NLR Immune Receptor RGA5". The Plant Cell. 29 (1): 156–168. doi:10.1105/tpc.16.00435. PMC 5304345. PMID 28087830.
- ↑ Shao F, Golstein C, Ade J, Stoutemyer M, Dixon JE, Innes RW (August 2003). "Cleavage of Arabidopsis PBS1 by a bacterial type III effector". Science. 301 (5637): 1230–3. Bibcode:2003Sci...301.1230S. doi:10.1126/science.1085671. PMID 12947197. S2CID 6418384.
- ↑ Adachi H, Kamoun S, Maqbool A (May 2019). "A resistosome-activated 'death switch'". Nature Plants. 5 (5): 457–458. doi:10.1038/s41477-019-0425-9. PMID 31036914. S2CID 139104570.
- ↑ van Wersch S, Li X (August 2019). "Stronger When Together: Clustering of Plant NLR Disease resistance Genes". Trends in Plant Science. 24 (8): 688–699. doi:10.1016/j.tplants.2019.05.005. PMID 31266697. S2CID 195787161.
- ↑ Césari S, Kanzaki H, Fujiwara T, Bernoux M, Chalvon V, Kawano Y, et al. (September 2014). "The NB-LRR proteins RGA4 and RGA5 interact functionally and physically to confer disease resistance". The EMBO Journal. 33 (17): 1941–59. doi:10.15252/embj.201487923. PMC 4195788. PMID 25024433.
- 1 2 3 Wu CH, Abd-El-Haliem A, Bozkurt TO, Belhaj K, Terauchi R, Vossen JH, Kamoun S (July 2017). "NLR network mediates immunity to diverse plant pathogens". Proceedings of the National Academy of Sciences of the United States of America. 114 (30): 8113–8118. Bibcode:2017PNAS..114.8113W. doi:10.1073/pnas.1702041114. PMC 5544293. PMID 28698366.
- ↑ Adachi H, Contreras MP, Harant A, Wu CH, Derevnina L, Sakai T, et al. (November 2019). "An N-terminal motif in NLR immune receptors is functionally conserved across distantly related plant species". eLife. 8. doi:10.7554/eLife.49956. PMC 6944444. PMID 31774397.
- ↑ Lai Y, Eulgem T (May 2018). "Transcript-level expression control of plant NLR genes". Molecular Plant Pathology. 19 (5): 1267–1281. doi:10.1111/mpp.12607. PMC 6638128. PMID 28834153.
- ↑ Azevedo C, Betsuyaku S, Peart J, Takahashi A, Noël L, Sadanandom A, et al. (May 2006). "Role of SGT1 in resistance protein accumulation in plant immunity". The EMBO Journal. 25 (9): 2007–16. doi:10.1038/sj.emboj.7601084. PMC 1456927. PMID 16619029.
- ↑ Whitham S, McCormick S, Baker B (August 1996). "The N gene of tobacco confers resistance to tobacco mosaic virus in transgenic tomato". Proceedings of the National Academy of Sciences of the United States of America. 93 (16): 8776–81. Bibcode:1996PNAS...93.8776W. doi:10.1073/pnas.93.16.8776. PMC 38750. PMID 8710948.
- ↑ Bieri S, Mauch S, Shen QH, Peart J, Devoto A, Casais C, et al. (December 2004). "RAR1 positively controls steady state levels of barley MLA resistance proteins and enables sufficient MLA6 accumulation for effective resistance". The Plant Cell. 16 (12): 3480–95. doi:10.1105/tpc.104.026682. PMC 535887. PMID 15548741.
- ↑ Jones JD, Vance RE, Dangl JL (December 2016). "Intracellular innate immune surveillance devices in plants and animals". Science. 354 (6316): aaf6395. doi:10.1126/science.aaf6395. PMID 27934708.
- ↑ Liu Y, Ren D, Pike S, Pallardy S, Gassmann W, Zhang S (September 2007). "Chloroplast-generated reactive oxygen species are involved in hypersensitive response-like cell death mediated by a mitogen-activated protein kinase cascade". The Plant Journal. 51 (6): 941–54. doi:10.1111/j.1365-313X.2007.03191.x. PMID 17651371.
- ↑ Koyanagi T, Matsumura K, Kuroda S, Tanizawa K (April 2000). "Molecular cloning and heterologous expression of pea seedling copper amine oxidase". Bioscience, Biotechnology, and Biochemistry. 64 (4): 717–22. doi:10.1271/bbb.64.717. PMID 10830482.
- ↑ Abramovitch RB, Martin GB (August 2004). "Strategies used by bacterial pathogens to suppress plant defenses". Current Opinion in Plant Biology. 7 (4): 356–64. doi:10.1016/j.pbi.2004.05.002. PMID 15231256.
- ↑ Grant M, Lamb C (August 2006). "Systemic immunity". Current Opinion in Plant Biology. 9 (4): 414–20. doi:10.1016/j.pbi.2006.05.013. PMID 16753329.
- ↑ Tran DT, Chung EH, Habring-Müller A, Demar M, Schwab R, Dangl JL, et al. (April 2017). "Activation of a Plant NLR Complex through Heteromeric Association with an Autoimmune Risk Variant of Another NLR". Current Biology. 27 (8): 1148–1160. doi:10.1016/j.cub.2017.03.018. PMC 5405217. PMID 28416116.
- ↑ Phadnis N, Malik HS (December 2014). "Speciation via autoimmunity: a dangerous mix". Cell. 159 (6): 1247–9. doi:10.1016/j.cell.2014.11.028. PMID 25480288.
- 1 2 Maekawa T, Kufer TA, Schulze-Lefert P (August 2011). "NLR functions in plant and animal immune systems: so far and yet so close". Nature Immunology. 12 (9): 817–26. doi:10.1038/ni.2083. PMID 21852785. S2CID 205364432.
- ↑ Burdett H, Kobe B, Anderson PA (July 2019). "Animal NLRs continue to inform plant NLR structure and function" (PDF). Archives of Biochemistry and Biophysics. 670: 58–68. doi:10.1016/j.abb.2019.05.001. PMID 31071301. S2CID 149446493.
- ↑ Liu X, Zhang Z, Ruan J, Pan Y, Magupalli VG, Wu H, Lieberman J (July 2016). "Inflammasome-activated gasdermin D causes pyroptosis by forming membrane pores". Nature. 535 (7610): 153–8. Bibcode:2016Natur.535..153L. doi:10.1038/nature18629. PMC 5539988. PMID 27383986.
- ↑ Wang J, Hu M, Wang J, Qi J, Han Z, Wang G, et al. (April 2019). "Reconstitution and structure of a plant NLR resistosome conferring immunity". Science. 364 (6435): eaav5870. doi:10.1126/science.aav5870. PMID 30948527. S2CID 96434803.
- ↑ Dickman M, Williams B, Li Y, de Figueiredo P, Wolpert T (October 2017). "Reassessing apoptosis in plants". Nature Plants. 3 (10): 773–779. doi:10.1038/s41477-017-0020-x. PMID 28947814. S2CID 3290201.