The RNA integrity number (RIN) is an algorithm for assigning integrity values to RNA measurements.
The integrity of RNA is a major concern for gene expression studies and traditionally has been evaluated using the 28S to 18S rRNA ratio, a method that has been shown to be inconsistent.[1] This inconsistency arises because subjective, human interpretation is necessary to compare the 28S and 18S gel images. The RIN algorithm was devised to overcome this issue. The RIN algorithm is applied to electrophoretic RNA measurements, typically obtained using capillary gel electrophoresis, and based on a combination of different features that contribute information about the RNA integrity to provide a more universal measure. RIN has been demonstrated to be robust and reproducible in studies comparing it to other RNA integrity calculation algorithms, cementing its position as a preferred method of determining the quality of RNA to be analyzed.[2]
A major criticism to RIN is when using with plants or in studies of eukaryotic-prokaryotic cells interactions. The RIN algorithm is unable to differentiate eukaryotic/prokaryotic/chloroplastic ribosomal RNA, creating serious quality index underestimation in such situations.
Terminology
Electrophoresis is the process of separating nucleic acid species based on their length by applying an electric field to them. As nucleic acids are negatively charged, they are pushed by an electric field through a matrix, usually an agarose gel, with the smaller molecules being pushed farther, faster.[3] Capillary electrophoresis is a technique whereby small amounts of a nucleic acid sample can be run on a gel in a very thin tube. There is a detector in the machine that can tell when nucleic acid samples pass through a specific point in the tube, with smaller samples passing through first. This can produce an electropherogram such as the one in Figure 1, where length is related to time at which the samples pass the detector.
A marker is a sample of known size run along with the sample so that the actual size of the rest of the sample can be known by comparing their running distance/time to be relative to this marker.
RNA is a biological macromolecule made of sugars and nitrogenous bases that plays a number of crucial roles in all living cells. There are several subtypes of RNA, with the most prominent in the cell being tRNA (transfer RNA), rRNA (ribosomal RNA), and mRNA (messenger RNA). All three of these are involved in the process of translation, with the most prominent species (~85%) of cellular RNA being rRNA. As a result, this is the most immediately visible species when RNA is analyzed via electrophoresis and is thus used for determining RNA quality (see Computation, below). rRNA comes in various sizes, with those in mammals belonging to the sizes 5S, 18S, and 28S. The 28S and 5S rRNAs form the large subunit and the 18S forms the small subunit of the ribosome, the molecular machinery responsible for synthesizing proteins.[4]
Applications
RNases are ubiquitous and can often contaminate and subsequently degrade RNA samples in the laboratory, so RNA integrity can very easily be compromised, leading to a number of laboratory techniques designed to eliminate their impact.[5][6] However, these methods are not fool-proof, and so samples can still be degraded, necessitating a method of measuring RNA integrity to ensure the trustworthiness and reproducibility of molecular assays, as RNA integrity is critical for proper results in gene expression studies, such as microarray analysis, Northern blots, or quantitative real-time PCR (qPCR).[7][8] RNA that has been degraded has a direct impact on calculated expression levels, often leading to significantly decreased apparent expression.[9]
qPCR and similar techniques are very expensive, taking a good deal of both time and money, so continuing research being undertaken to decrease the cost while maintaining qPCR's accuracy and reproducibility for gene expression and other applications.[10] RIN assessment allows a scientist to evaluate an experiment's trustworthiness and reproducibility before incurring substantial costs in performing the gene expression studies.
RIN is a standard method of measuring RNA integrity and can be used to evaluate the quality of RNA produced by new RNA isolation techniques.[11]
Development
As RNA integrity has long been known to be a problem in molecular biology studies, there are a few methods that have been used historically to determine the integrity of RNA. The most popular has long been agarose gel electrophoresis with ethidium bromide staining, allowing one to visualize the bands from the rRNA peaks. The height of the 28S and 18S bands can be compared to each other, with a 2:1 ratio indicating non-degraded RNA.[1] While this method is very cheap and easy, there are several issues with this method, primarily its subjectivity, leading to inconsistent, non-standardized RNA quality assessments, and the large amounts of RNA that are needed to visualize it on an agarose gel, which can be problematic if there is not much RNA to work with.[1][12] There are also a number of different problems that can arise from agarose gel electrophoresis, such as poor loading, uneven running, and uneven staining that lead to increased variability in the accuracy of using agarose gel electrophoresis to determine RNA integrity.[13]
The RNA Integrity Number was developed by Agilent Technologies in 2005.[1] The algorithm was generated by taking hundreds of samples and having specialists manually assign them all a value of 1 to 10 based on their integrity, with 10 being the highest. Adaptive learning tools using a Bayesian learning technique were used to generate an algorithm that could predict the RIN, predominantly by using the features listed below under "Computation".[1][14] This allows for all Agilent software to produce the same RIN for a given RNA sample, standardizing the measurement and making it much less subjective than earlier methods.
Computation
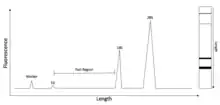

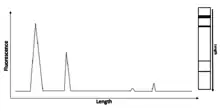
RIN for a sample is computed using several characteristics of an RNA electropherogram trace, with the first two listed below being most significant. RIN assigns an electropherogram a value of 1 to 10, with 10 being the least degraded. All the following descriptions apply to mammalian RNA because RNAs in other species have different rRNA sizes:[1] The total RNA ratio is calculated by taking the ratio of the area under the 18S and 28S rRNA peaks to the total area under the graph, a large number here is desired, indicating much of the rRNA is still at these sizes and thus little to no degradation has occurred. An ideal ratio can be seen in figure 1, where almost all of the RNA is in the 18S and 28S RNA peaks.
For the height of 28S peak, a large value is desired. 28S, the most prominent rRNA species, is used in RIN calculation as it is typically degraded more quickly than 18S rRNA, and so measuring its peak height allows for detection of the early stages of degradation. Again, this is seen in figure 1, where the 28S peak is the largest, and so this is good.
The fast region is the area between the 18S and 5S rRNA peaks on an electropherogram. Initially, as the fast area ratio value increases, it indicates degradation of the 18S and 28S rRNA to an intermediate size, though the ratio subsequently decreases as RNA degrades further, to even smaller sizes. Thus, a low value doesn't necessarily indicate either good or bad RNA integrity.
A small marker height is desired, indicating only small amounts of RNA have been degraded and proceeded to the smallest lengths, indicated by the short marker. If a large number is found here, that indicates that large amounts of the rRNAs have been degraded to small pieces that would be found closer to this marker. This situation can be seen in the 'poor quality' RNA electropherogram found in figure 2, where the height of the peak over the marker (far left) is very large, so the RNA has been greatly degraded. In prokaryotic samples, the algorithm is somewhat different, but the Agilent 2100 Bioanalyzer Expert software is able to calculate RIN for prokaryotic samples now as well.[15] The difference likely arises from the fact that, while mammalian samples have 28S and 18S ribosomal RNAs as their predominant species, prokaryotic RNAs have the sizes shifted slightly smaller, to 23S and 16S, so the algorithm must be shifted to accommodate that. Another crucial fact about calculating prokaryotic RNA integrity numbers is that RIN has not been validated to the extent that it has for eukaryotic RNA.[15] It has been shown that higher RIN values correlate with better downstream results in eukaryotes, but this hasn't been done as extensively for prokaryotes, so it may mean less in prokaryotes.
These electropherograms for calculating RIN are done using the Agilent Bioanalyzer machine, which is capable of performing electrophoresis and generating the electropherograms.[14] The Agilent 2100 software is uniquely able to perform the RIN software, as the exact algorithm is proprietary, so there are additional important RNA electropherogram features that are used in its calculation that are not publicly available.
References
- 1 2 3 4 5 6 Schroeder, Andreas; Mueller, Odilo; Stocker, Susanne; Salowsky, Ruediger; Leiber, Michael; Gassmann, Marcus; Lightfoot, Samar; Menzel, Wolfram; Granzow, Martin; Ragg, Thomas (2006). "The RIN: an RNA integrity number for assigning integrity values to RNA measurements". BMC Molecular Biology. 7 (1): 3. doi:10.1186/1471-2199-7-3. PMC 1413964. PMID 16448564.
- ↑ Imbeaud, Sandrine; Graudens, Esther; Boulanger, Virginie; Barlet, Xavier; Zaborski, Patrick; Eveno, Eric; Mueller, Odilo; Schroeder, Andreas; Auffray, Charles (2005-01-01). "Towards standardization of RNA quality assessment using user-independent classifiers of microcapillary electrophoresis traces". Nucleic Acids Research. 33 (6): e56. doi:10.1093/nar/gni054. ISSN 0305-1048. PMC 1072807. PMID 15800207.
- ↑ Watson, James D.; Baker, Tania A.; Bell, Stephen P.; Gann, Alexander; Levine, Michael; Losick, Richard (2014). Molecular Biology of the Gene: Seventh Edition. Glenview, IL: Pearson Education. ISBN 978-0-321-85149-9.
- ↑ Khatter, Heena; Myasnikov, Alexander G.; Natchiar, S. Kundhavai; Klaholz, Bruno P. (2015). "Structure of the human 80S ribosome". Nature. 520 (7549): 640–645. Bibcode:2015Natur.520..640K. doi:10.1038/nature14427. PMID 25901680. S2CID 4459012.
- ↑ "Avoiding Ribonuclease Contamination | NEB". www.neb.com. Retrieved 2016-04-10.
- ↑ "The Basics: RNase Control". www.thermofisher.com. Retrieved 2016-04-10.
- ↑ Bustin, Stephen A.; Benes, Vladimir; Garson, Jeremy A.; Hellemans, Jan; Huggett, Jim; Kubista, Mikael; Mueller, Reinhold; Nolan, Tania; Pfaffl, Michael W. (2009-04-01). "The MIQE Guidelines: Minimum Information for Publication of Quantitative Real-Time PCR Experiments". Clinical Chemistry. 55 (4): 611–622. doi:10.1373/clinchem.2008.112797. ISSN 0009-9147. PMID 19246619.
- ↑ Fleige, Simone; Pfaffl, Michael W. (2006). "RNA integrity and the effect on the real-time QRT-PCR performance". Molecular Aspects of Medicine. 27 (2–3): 126–139. doi:10.1016/j.mam.2005.12.003. PMID 16469371.
- ↑ Taylor, Sean C.; Mrkusich, Eli M. (2014). "The State of RT-Quantitative PCR: Firsthand Observations of Implementation of Minimum Information for the Publication of Quantitative Real-Time PCR Experiments (MIQE)". Journal of Molecular Microbiology and Biotechnology. 24 (1): 46–52. doi:10.1159/000356189. PMID 24296827.
- ↑ Reitschuler, Christoph; Lins, Philipp; Illmer, Paul (2014). "Primer evaluation and adaption for cost-efficient SYBR Green-based QPCR and its applicability for specific quantification of methanogens". World Journal of Microbiology and Biotechnology. 30 (1): 293–304. doi:10.1007/s11274-013-1450-x. PMID 23918633. S2CID 36790110.
- ↑ Livingston, Whitney S.; Rusch, Heather L.; Nersesian, Paula V.; Baxter, Tristin; Mysliwiec, Vincent; Gill, Jessica M. (2015-01-01). "Improved sleep in military personnel is associated with changes in the expression of inflammatory genes and improvement in depression symptoms". Frontiers in Psychiatry. 6: 59. doi:10.3389/fpsyt.2015.00059. PMC 4415307. PMID 25983695.
- ↑ Taylor, Sean; Wakem, Michael; Dijkman, Greg; Alsarraj, Marwan; Nguyen, Marie (2010-04-01). "A practical approach to RT-qPCR—Publishing data that conform to the MIQE guidelines". Methods. The ongoing Evolution of qPCR. 50 (4): S1–S5. doi:10.1016/j.ymeth.2010.01.005. PMID 20215014.
- ↑ "Advancing the quality control methodology to assess isolated total RNA and generated fragmented cRNA", Agilent Application Note, Publication Number 5988-9861EN, 2003.
- 1 2 "RNA Integrity Number (RIN) – Standardization of RNA Quality Control", Agilent Application Note, Publication Number 5989-1165EN, 2016.
- 1 2 Agilent 2100 Bioanalyzer: 2100 Expert User’s Guide. Waldbronn, Germany: Agilent Technologies (2005).
External links
- RIN information from Agilent Technologies
- RIN article in BMC Molecular Biology
- Gene expression summary from Nature to help show why we need RIN
- RIN explanation from Agilent Technologies, with several examples of electropherograms at various RIN values
- Gel electrophoresis simulation from the University of Utah to help visualize how the electropherograms are produced