| Ets-domain | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
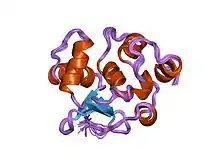 Structure of Ets-1 DNA binding autoinhibition.[1] | |||||||||||
| Identifiers | |||||||||||
| Symbol | Ets | ||||||||||
| Pfam | PF00178 | ||||||||||
| Pfam clan | CL0123 | ||||||||||
| InterPro | IPR000418 | ||||||||||
| SMART | SM00413 | ||||||||||
| PROSITE | PDOC00374 | ||||||||||
| SCOP2 | 1r36 / SCOPe / SUPFAM | ||||||||||
| |||||||||||
In the field of molecular biology, the ETS (E26 transformation-specific[2] or E-twenty-six. (Erythroblast Transformation Specific)[3]) family is one of the largest families of transcription factors and is unique to animals. There are 29 genes in humans, 28 in the mouse, 10 in Caenorhabditis elegans and 9 in Drosophila. The founding member of this family was identified as a gene transduced by the leukemia virus, E26. The members of the family have been implicated in the development of different tissues as well as cancer progression.
Subfamilies
The ETS (Erythroblast Transformation Specific)family is divided into 12 subfamilies, which are listed below:[4]
| Subfamily | Mammalian family members | Invertebrate orthologs |
|---|---|---|
| ELF | ELF1, ELF2 (NERF), ELF4 (MEF) | |
| ELG | GABPα | ELG |
| ERG | ERG, FLI1, FEV | |
| ERF | ERF (PE2), ETV3 (PE1) | |
| ESE | ELF3 (ESE1/ESX), ELF5 (ESE2), ESE3 (EHF) | |
| ETS | ETS1, ETS2 | POINTED |
| PDEF | SPDEF (PDEF/PSE) | |
| PEA3 | ETV4 (PEA3/E1AF), ETV5 (ERM), ETV1 (ER81) | |
| ER71 | ETV2 (ER71) | |
| SPI | SPI1 (PU.1), SPIB, SPIC | |
| TCF | ELK1, ELK4 (SAP1), ELK3 (NET/SAP2) | LIN |
| TEL | ETV6 (TEL), ETV7 (TEL2) | YAN |
Structure
All ETS (Erythroblast Transformation Specific) family members are identified through a highly conserved DNA binding domain, the ETS domain, which is a winged helix-turn-helix structure that binds to DNA sites with a central GGA(A/T) DNA sequence. As well as DNA-binding functions, evidence suggests that the ETS domain is also involved in protein-protein interactions. There is limited similarity outside the ETS DNA binding domain.
Other domains are also present and vary from ETS member to ETS member, including the Pointed domain, a subclass of the SAM domain family.
Function
The ETS family is present throughout the body and is involved in a wide variety of functions including the regulation of cellular differentiation, cell cycle control, cell migration, cell proliferation, apoptosis (programmed cell death) and angiogenesis.
Multiple ETS factors have been found to be associated with cancer, such as through gene fusion. For example, the ERG ETS transcription factor is fused to the EWS gene, resulting in a condition called Ewing's sarcoma.[5] The fusion of TEL to the JAK2 protein results in early pre-B acute lymphoid leukaemia.[6] ERG and ETV1 are known gene fusions found in prostate cancer.[7]
In addition, ETS factors, e.g. the vertebrate Etv1 and the invertebrate Ast-1, have been shown to be important players in the specification and differentiation of dopaminergic neurons in both C. elegans and olfactory bulbs of mice.[8]
Mode of action
Amongst members of the ETS family, there is extensive conservation in the DNA-binding ETS domain and, therefore, a lot of redundancy in DNA binding. It is thought that interactions with other proteins (eg: Modulator of the activity of Ets called Mae) is one way in which specific binding to DNA is achieved. Transcription factor Ets are a site of signalling convergence.[9] ETS factors act as transcriptional repressors, transcriptional activators, or both.[10]
References
- ↑ Lee GM, Donaldson LW, Pufall MA, et al. (February 2005). "The structural and dynamic basis of Ets-1 DNA binding autoinhibition". J. Biol. Chem. 280 (8): 7088–99. doi:10.1074/jbc.M410722200. PMID 15591056.
- ↑ Nunn, M. F.; Seeburg, P. H.; Moscovici, C.; Duesberg, P. H. (1983). "Tripartite structure of the avian erythroblastosis virus E26 transforming gene". Nature. 306 (5941): 391–395. Bibcode:1983Natur.306..391N. doi:10.1038/306391a0. PMID 6316155. S2CID 4302399.
- ↑ Leprince, D.; Gegonne, A.; Coll, J.; De Taisne, C.; Schneeberger, A.; Lagrou, C.; Stehelin, D. (1983). "A putative second cell-derived oncogene of the avian leukaemia retrovirus E26". Nature. 306 (5941): 395–397. Bibcode:1983Natur.306..395L. doi:10.1038/306395a0. PMID 6316156. S2CID 4318034.
- ↑ Gutierrez-Hartman A, Duval DL, Bradford AP (2007). "ETS transcription factors in endocrine systems". Trends Endocrinol Metab. 18 (4): 150–8. doi:10.1016/j.tem.2007.03.002. PMID 17387021. S2CID 24617218.
- ↑ Ida K, Kobayashi S, Taki T, Hanada R, Bessho F, Yamamori S, Sugimoto T, Ohki M, Hayashi Y (1995). "EWS-FLI-1 and EWS-ERG chimeric mRNAs in Ewing's sarcoma and primitive neuroectodermal tumor". Int J Cancer. 63 (4): 500–4. doi:10.1002/ijc.2910630407. PMID 7591257. S2CID 24841690.
- ↑ Peeters P, Raynaud SD, Cools J, Wlodarska I, Grosgeorge J, Philip P, Monpoux F, Van Rompaey L, Baens M, Van den Berghe H, Marynen P (1997). "Fusion of TEL, the ETS-variant gene 6 (ETV6), to the receptor-associated kinase JAK2 as a result of t(9;12) in a lymphoid and t(9;15;12) in a myeloid leukemia". Blood. 90 (7): 2535–40. doi:10.1182/blood.V90.7.2535. PMID 9326218.
- ↑ Tomlins SA, Rhodes DR, Perner S, Dhanasekaran SM, Mehra R, Sun XW, Varambally S, Cao X, Tchinda J, Kuefer R, Lee C, Montie JE, Shah RB, Pienta KJ, Rubin MA, Chinnaiyan AM (October 2005). "Recurrent fusion of TMPRSS2 and ETS transcription factor genes in prostate cancer". Science. 310 (5748): 644–8. Bibcode:2005Sci...310..644T. doi:10.1126/science.1117679. PMID 16254181. S2CID 85788789.
- ↑ Flames N, Hobert O (2009). "Gene regulatory logic of dopaminergic neuron differentiation". Nature. 458 (7240): 885–890. doi:10.1038/nature07929. PMC 2671564. PMID 19287374.
- ↑ Verger A, Duterque-Coquillaud M (2002). "When Ets transcription factors meet their partners". BioEssays. 24 (4): 362–70. doi:10.1002/bies.10068. PMID 11948622.
- ↑ Sharrocks AD (2001). "The ETS-domain transcription factor family". Nat Rev Mol Cell Biol. 2 (11): 827–37. doi:10.1038/35099076. PMID 11715049. S2CID 5407789.
Further reading
- Blair DG, Athanasiou M (2000). "Ets and retroviruses - transduction and activation of members of the Ets oncogene family in viral oncogenesis". Oncogene. 19 (55): 6472–81. doi:10.1038/sj.onc.1204046. PMID 11175363.
- Li R, Pei H, Watson DK (2000). "Regulation of Ets function by protein - protein interactions". Oncogene. 19 (55): 6514–23. doi:10.1038/sj.onc.1204035. PMID 11175367.
- Mimeault M (2000). "Structure-function studies of ETS transcription factors". Crit Rev Oncog. 11 (3–4): 227–53. doi:10.1615/critrevoncog.v11.i34.20. PMID 11358268.
- Oikawa T, Yamada T (2003). "Molecular biology of the Ets family of transcription factors". Gene. 303: 11–34. doi:10.1016/S0378-1119(02)01156-3. PMID 12559563.
- Sementchenko VI, Watson DK (2000). "Ets target genes: past, present and future". Oncogene. 19 (55): 6533–48. doi:10.1038/sj.onc.1204034. PMID 11175369.
- Verger A, Duterque-Coquillaud M (2002). "When Ets transcription factors meet their partners". BioEssays. 24 (4): 362–70. doi:10.1002/bies.10068. PMID 11948622.
- Nunn M, Seeburg P, Moscovici C, Duesberg P (1983). "Tripartite structure of the avian erythroblastosis virus E26 transforming gene". Nature. 306 (5941): 391–95. Bibcode:1983Natur.306..391N. doi:10.1038/306391a0. PMID 6316155. S2CID 4302399.